23 July 2020: Database Analysis
Identification of Biomarkers for Sarcoidosis and Tuberculosis of the Lung Using Systematic and Integrated Analysis
Min Zhao1ABCEF, Xin Di1BCE, Xin Jin2CDE, Chang Tian1BF, Shan Cong1CF, Jiaying Liu1BF, Ke Wang1ABDEG*DOI: 10.12659/MSM.925438
Med Sci Monit 2020; 26:e925438
Abstract
BACKGROUND: Sarcoidosis (SARC) is a multisystem inflammatory disease of unknown etiology and pulmonary tuberculosis (PTB) is caused by Mycobacterium tuberculosis. Both of these diseases affect lungs and lymph nodes and share similar clinical manifestations. However, the underlying mechanisms for the similarities and differences in genetic characteristics of SARC and PTB remain unclear.
MATERIAL AND METHODS: Three datasets (GSE16538, GSE20050, and GSE19314) were retrieved from the Gene Expression Omnibus (GEO) database. Differentially expressed genes (DEGs) in SARC and PTB were identified using GEO2R online analyzer and Venn diagram software. Functional enrichment analysis was performed using Database for Annotation, Visualization and Integrated Discovery (DAVID) and R packages. Two protein–protein interaction (PPI) networks were constructed using Search Tool for the Retrieval of Interacting Genes database, and module analysis was performed using Cytoscape. Hub genes were identified using area under the receiver operating characteristic curve analysis.
RESULTS: We identified 228 DEGs, including 56 common SARC-PTB DEGs (enriched in interferon-gamma-mediated signaling, response to gamma radiation, and immune response) and 172 SARC-only DEGs (enriched in immune response, cellular calcium ion homeostasis, and dendritic cell chemotaxis). Potential biomarkers for SARC included CBX5, BCL11B, and GPR18.
CONCLUSIONS: We identified potential biomarkers that can be used as candidates for diagnosis and/or treatment of patients with SARC.
Keywords: Biological Markers, Gene Expression Profiling, sarcoidosis, Tuberculosis, Biomarkers, Tumor, Chromobox Protein Homolog 5, Computational Biology, Databases, Genetic, Gene Expression Regulation, gene ontology, Gene Regulatory Networks, Lung, Protein Interaction Mapping, Protein Interaction Maps, Software
Background
Sarcoidosis (SARC) is a multisystem inflammatory disease of unknown etiology, and primarily affects the lungs and thoracic lymph nodes and manifests with noncaseating epithelioid granulomas (aggregates of lymphocytes, macrophages, epithelioid, and giant cells) [1]. Granulomatous diseases result from the continued presentation of a poorly degradable antigen [2]. Among these, the infectious agents
Microarray technology provides an all-in-one system to simultaneously scan hybridization signals from tens of thousands of gene probes on a chip and quantitatively analyze the transcriptome of samples [7]. Transcriptomic analysis can be used to understand various aspects of SARC and PTB. These profiles are stored in genomic databases that need to be analyzed to understand the characteristics of diseases, such as SARC and PTB. This enables the identification of potential candidates that can be used for diagnosis and treatment. In this study, we compared the expression profiles of lung tissues from patients and healthy individuals to identify disease signatures and understand the pathogenesis of SARC. We used the GSE16538, GSE20050, and GSE19314 expression microarray data from the Gene Expression Omnibus (GEO) database to identify differentially expressed genes (DEGs) and their interaction patterns using a protein–protein interaction (PPI) network. DEG functions were determined using Gene Ontology (GO) and pathway enrichment analysis. A number of genes were putative (novel) candidates for use as diagnostic, therapeutic, and prognostic markers for SARC. This will help develop new array-based diagnostic tools to distinguish patients from healthy individuals and differentiate between diseases of similar pathology. Differential biomarker profiles can also provide insight into the common biological processes affected by similar diseases, such as SARC and PTB.
Material and Methods
DATA DOWNLOAD AND PRE-PROCESSING:
NCBI-GEO is a free public database of microarray/genomic data repository. The gene expression profiles in lung tissues from SARC patients, lung tissues from PTB patients, and peripheral blood from SARC patients were obtained from the GSE16538, GSE20050, and GSE19314 datasets, respectively. Microarray data of GSE16538 and GSE19314 were obtained using GPL570 Platforms ([HG-U133_Plus_2] Affymetrix Human Genome U133 Plus 2.0 Array) that included 6 tissue samples from patients with SARC and healthy individuals each, and 37 and 20 blood samples from patients with SARC and healthy individuals, respectively. Microarray data from GSE20050 was obtained using the GPL1352 platform ([U133_X3P] Affymetrix Human X3P Array) comprising 3 PTB and 2 healthy lung tissue samples (Table 1).
DATA PROCESSING AND IDENTIFICATION OF DEGS:
DEGs in the lung samples from patients with SARC and PTB and healthy individuals were identified using the GEO2R online tools with fold change (|logFC|) >1 and adjusted
GO AND PATHWAY ENRICHMENT ANALYSIS:
GO is commonly used to determine the function(s) of genes during the analysis of high throughput transcriptomic or genomic data. Kyoto Encyclopedia of Genes and Genomes (KEGG) is a collection of databases associated with genomes, diseases, biological pathways, drugs, and chemical materials. DAVID (Database for Annotation, Visualization and Integrated Discovery) is an online bioinformatics tool that is used to identify the functions of genes or proteins. We used DAVID to determine the functions of DEGs in the biological process (BP), cellular component (CC), molecular function (MF), and pathways (
PPI NETWORKS AND MODULE ANALYSIS:
PPI networks help understand the molecular mechanisms underlying biological progresses. PPI networks was generated by Search Tool for the Retrieval of Interacting Genes (STRING) (
IDENTIFICATION OF HUB GENE(S):
The hub genes in the candidate modules were identified using Cytoscape plugin cytoHubba (top 10 nodes ranked by MCC, MNC, Radiality, Stress, Betweenness, BottleNeck, Closeness, ClusteringCoefficient, Degree, DMNC, EcCentricity, and EPC). We used the expression data of GSE19314 database to generate the receiver operating characteristic (ROC) curves of the hub genes by R/Bioconductor. The area under the ROC curve (AUC) for these genes were calculated to test their potential in diagnosing SARC using. AUC >0.5 was considered statistically significant.
Results
IDENTIFICATION OF DEGS IN SARC AND PTB:
We identified 228 and 2913 DEGs from the GSE16538 and GSE20050 databases, respectively. Using Venn diagrams, we categorized the DEGs related to SARC that were shared with PTB or characteristic of SARC (SARC-PTB common and SARC-only DEGs). There were 56 SARC-PTB common DEGs, including 1 downregulated gene (logFC <0) and 55 upregulated genes (logFC >0; Table 2). We identified 172 SARC-only DEGs, including 30 downregulated genes and 142 upregulated genes (Table 3, Figure 1).
GO AND KEGG ANALYSIS:
The 56 SARC-PTB common DEGs were analyzed by DAVID followed by the R package to analyze their biological functions and associated pathways. DEGs were enriched in BP involved in interferon-gamma-mediated signaling pathway, response to gamma radiation, immune response, response to hydrogen peroxide, defense response, response to nutrient, signal transduction, apoptotic process, regulation of interferon-gamma-mediated signaling pathway, and intrinsic apoptotic signaling pathway in response to oxidative stress; in the CC comprising the membrane; and MF in protein homodimerization activity, signal transducer activity, and anion transmembrane transporter activity (Table 4). Table 5 shows the KEGG analysis highlighting the enrichments of DEGs in pathways related to cancer and chemokine signaling pathway (P<0.05).
The 172 SARC-only upregulated DEGs were enriched in BP involved in immune response, cellular calcium ion homeostasis, dendritic cell chemotaxis, chemokine-mediated signaling pathway, positive regulation of Rho protein signal transduction, signal transduction, cell maturation, RNA processing, regulation of transcription from RNA polymerase II promoter, negative regulation of chemokine-mediated signaling pathway, regulation of T cell differentiation in thymus, and response to virus; the CC of neuronal ribonucleoprotein granule and histone deacetylase complex; and in MF involved in metal ion binding, nucleic acid binding, RNA polymerase II core promoter proximal region sequence-specific DNA binding, ATP binding, protein binding, chemokine activity, and manganese ion binding. The downregulated DEGs were enriched in BP involved in response to vitamin D, CC including the extracellular space, extracellular exosome, and endosome membrane, and MF in transporter activity (Figure 2A–2F). Table 6 shows the KEGG analysis highlighting that upregulated DEGs were particularly enriched in chemokine signaling pathway, microRNAs in cancer, and cytokine–cytokine receptor interaction; no KEGG pathway was associated with downregulated DEGs (P<0.05).
PPI NETWORK AND MODULE ANALYSIS:
PPI networks reflect the spatiotemporal relationship of macromolecules within the cell that helps understand the molecular mechanisms involved in physiological and pathological processes. To explore the molecular mechanism(s) underlying SARC progression, we used the STRING database to construct the PPI network. An interaction score of 0.4 (highest confidence) was set as threshold. The PPI network for SARC-PTB common DEGs consisted of 31 nodes and 31 edges (Figure 3A) and that of SARC-only DEGs comprised 95 nodes with 139 edges (Figure 4A). Subsequently, we identified the key modules using the MCODE plugin. We detected one functional cluster module and related genes in the PPI network of SARC-PTB common DEGs (Figure 3B). KEGG analysis of the genes in the module revealed the association with Toll-like receptor signaling pathway (hsa04620). A total of 3 functional clusters of modules and related genes were detected in the PPI network for SARC-only DEGs (Figure 4B–4D). KEGG analysis of the module genes revealed their association with cytokine-cytokine receptor interaction (hsa04060), chemokine signaling pathway (hsa04062), and RNA transport (hsa03013).
HUB GENE ANALYSIS:
The top 10 genes of each sub-plugin in the PPI network of SARC-only DEGs were identified by the cytoHubba plugin (Table 7). The 13 genes with the highest frequency in all sub-plugins were classified as hub genes, including IL7, DICER1, CCL19, CBX5, CDK6, CXCL11, IL15, GNG2, BCL11B, CCR2, CCL5, GPR18, and ZEB1; all these genes were upregulated. KEGG analysis of the hub genes showed their involvement in cytokine-cytokine receptor interaction (hsa04060), chemokine signaling pathway (hsa04062), and microRNAs in cancer (hsa05206). We used AUC analysis to evaluate their potential in diagnosing SARC using the expression data in the GSE19314 database. ROC-AUCs were higher than 0.50 for 12 of the genes (except CCL5). AUC >0.5 was considered statistically significant. The AUCs for CBX5, BCL11B, and GPR18 were 0.81, 0.91, and 0.87, respectively; these genes are of higher diagnostic potential since they were all higher than 0.80 (Figure 5).
Discussion
Multiple lines of evidence show that Mtb is a predisposing factor for SARC. SARC and PTB share histopathological characteristics, such as the presence of granulomas; thus, histopathological evidence can be used as the gold standard for diagnosis. Studying the gene expression profiles of tissues involved in pathogenesis is a direct tool for understanding the role of Mtb in SARC. Thus, we employed an integrated bioinformatics approach using 2 GEO datasets (GSE16538 and GSE20050) to identify 228 DEGs associated with SARC, including 56 SARC-PTB common DEGs and 172 SARC-only DEGs. There are more upregulated genes and less downregulated genes among these DEGs. Although the current etiology of sarcoidosis is not yet clear, sarcoidosis is closely related to the immune response. Excessive activation of the immune response to unknown inhaled antigens is considered to be one of the pathogenesis of sarcoidosis [8]. Most of the genes related to sarcoidosis that we have obtained are also related to the immune response, and most of DEGs are upregulated genes. We believe that the complex relationship of these immune-related upregulated genes may lead to excessive immune responses.
GO analysis revealed that the pathways enriched for SARC-PTB common DEGs were primarily associated with immune responses, especially interferon-gamma-mediated signaling. Unlike other chronic infectious diseases, Mtb promotes a powerful systemic immune response that damages local tissues and enhances the extended survival and spread of Mtb [9]. When the body is infected with an exogenous antigen, γ-interferon (IFN-γ) is released, following which local macrophages phagocytose, process and present antigens, and activate T cells. Subsequently, blood-derived lymphocytes and monocytes accumulate at the site of infection to form granulomas [10]. KEGG analysis demonstrated that SARC-PTB common DEGs were enriched in pathways related to cancer and the chemokine signaling pathway. This highlights the molecular interactions that are common to the pathogenesis of SARC and PTB. Some studies have shown a positive correlation between patient history of TB with increased risk for lung cancer and associated mortality. A Korean prospective cohort study showed the correlation between TB and risk of lung cancer in a high-income country [11]. Local chronic inflammation and oxidative stress are mechanisms that increase the risk of lung cancer in patients with a history of lung disease. Fibrosis is crucial in maintaining inflammation [12–16]. Similar studies have revealed a link between SARC and lung cancer [17–21]. The immunologic abnormalities in patients with SARC may be a result of the lack of immune response against tumors or oncogenic viruses. Conversely, SARC-related chronic inflammation may lead to the development of cancer [22].
GO and KEGG analysis of SARC-only DEGs showed enrichment in immune response, suggesting the involvement of immunity in the pathogenesis of SARC. These genes were also associated with BP related to viruses and vitamin D-related biological reactions. SARC may be caused by a variety of pathogens. In addition to Mtb and
In accordance with our KEGG analysis, among the 13 hub genes identified by the PPI network and modules analysis, IL7, CCR2, CCL19, IL15, and CXCL11 were associated with cytokine-cytokine receptor interaction. ROC curves demonstrated the diagnostic potential for these upregulated hub genes. IL7 is produced locally by intestinal epithelial and epithelial goblet cells and regulates intestinal mucosal lymphocytes. Knockout studies in mice have shown that this cytokine plays an essential role in lymphoid cell survival; it is important for B and T cell development. However, there is no report on the correlation between IL7 and SARC. CCR2 is expressed on the surface of macrophages, dendritic cells, and T lymphocytes and is a receptor for chemotactic proteins involved in inflammatory cell recruitment [32]. The research showed that CCR2 haplotype 2 is an independent factor that predisposes patients with SARC to Löfgren’s syndrome. CCR2 and its ligands have been shown to regulate cellular proliferation during granuloma formation in mice [33,34]. CCR2-deficient mice exhibit defective macrophage recruitment and impaired type-1 cytokine production. CCL19 is a CC-cytokine gene located on the p-arm of chromosome 9. Cytokines are a family of secreted proteins involved in immunoregulation and inflammation. Studies have shown that the overexpression of CCL19 in fluid from alveolar lavage obtained from patients with SARC promotes the development of alveolitis in the lungs of these patients by stimulating CD4+ lymphocyte recruitment [35,36]. IL15 is a cytokine that regulates T and natural killer cell activation and proliferation. IL15 and IL2 have similar biological functions. Tissues from patients with SARC possess a higher abundance of IL15+ cells as compared to that in asthmatic patients and healthy individuals. CXCL11 is a C-X-C motif chemokine that induces a chemotactic response in activated T cells and is the primary ligand for CXCR3. Studies shown that CXCL11 increases in the serum of patients with SARC as compared to that in healthy controls. Moreover, serum CXCL11 correlates with the severity of symptoms in patients with SARC, specifically organ involvement, respiratory symptoms, and abnormalities in pulmonary function [37]. Thus, CXCL11 may also be useful in identifying patients at risk for rapid pulmonary dysfunction.
ROC curve analysis showed high diagnostic potential (AUC >0.080) of CBX5, BCL11B, and GPR18. To the best of our knowledge, this is the first report to correlate these hub genes with SARC. CBX5 encodes a highly conserved non-histone member of the heterochromatin protein family. CBX5 is overexpressed in gastric cancer tissues and can be downregulated using vorinostat, indicating the oncogenic role of CBX5 in gastric cancer. MiR-758-3p targets CBX5 and inhibits the progression of gastric cancer [38]. Sarcoidosis is related to the occurrence of lung cancer, so it is possible that sarcoidosis-induced lung cancer is related to CBX5. BCL11B encodes a C2H2-type zinc finger protein that is closely related to BCL11A. A recent study based on HAART (the treatment regimen used for patients with HIV) showed that increased expression of BCL11B and its related chromatin modifiers contribute to the transcriptional silencing of HIV-1 and alternating BCL11B levels result in abnormal transcription and inflammation [39]. BCL11B is closely related to the immune response, so we speculate that there may be BCL11B participation in the pathogenesis of sarcoidosis, and more follow-up experiments are needed to demonstrate. GPR18, or N-arachidonyl glycine receptor (NAGly receptor), is a class A G-protein coupled receptor that, along with GPR119 and GPR55, comprises a novel cannabinoid receptor that consists of 331 amino acids. GPR18 is overexpressed in lymphoid tissues, such as the spleen and thymus, with moderate expression in the testes, ovaries, brain, and lungs. Current research, like in a rat model of inflammatory pain, an animal model of neuropathic pain (sciatic nerve injury), and an inflammatory pain model induced by intraplantar injection of complete Freund’s adjuvant (CFA), suggested GPR18 has an important role in mediating the analgesic properties of NAGly in different pain models [40]. GPR18 is highly expressed in lymphoid tissues and lungs, but the relationship between specific GPR18 and the respiratory system, including sarcoidosis, is rarely reported and requires further exploration. The molecular mechanism employed the potential hub gene candidates in patients with SARC need to be studied in detail.
In this study, we have identified a set of candidate biomarkers that can be exploited to study the pathogenesis, diagnosis, and prognosis of SARC.
Conclusions
In conclusion, we employed an integrated bioinformatics approach to identify the SARC-PTB common and SARC-only DEGs. A total of 13 hub genes from SARC-only DEGs were selected from the PPI network, among which 12 genes had the potential to be diagnostic biomarkers. CBX5, BCL11B, and CPR18 had the highest diagnostic potential. These genes are associated with cytokine-cytokine receptor interaction. However, this study had some limitations. This study was based on bioinformatics analysis of published data and lacks experimental verification. Thus, we could not determine the presence of a causal relationship between the differential expression of the hub genes and disease progression. Moreover, although we analyzed 3 GEO datasets, the sample size used was relatively small. This can lead to potential bias. Therefore, subsequent bioinformatics analyses and experimental verification with large sample sizes are necessary in the future to understand the molecular mechanism(s) employed by these novel hub genes in the pathogenesis of SARC and PTB. Furthermore, relevant models can also be constructed based on these genes to diagnose SARC and differentiate SARC from sputum negative PTB.
Figures
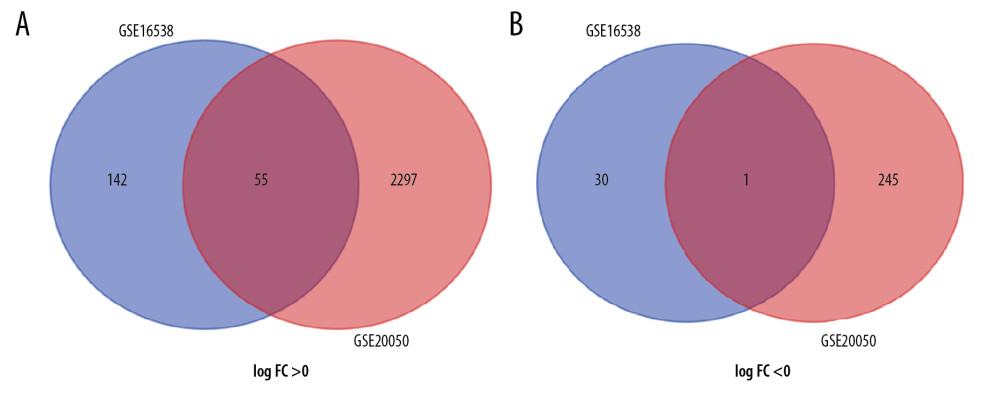 Figure 1. Identification of 56 common SARC-PTB DEGs in the 2 datasets (GSE16538 and GSE20050), and 172 SARC-only DEGs in GSE16538 through Venn diagrams software. Different color meant different datasets. (A) 55 DEGs were upregulated in the 2 datasets, 142 DEGs were in GSE16238 (logFC >0). (B) 1 DEG was downregulated in 2 datasets, 30 DEGs in GSE16238 (logFC <0). Different color meant different datasets. DEGs – differentially expressed genes; SARC – sarcoidosis; PTB – pulmonary tuberculosis.
Figure 1. Identification of 56 common SARC-PTB DEGs in the 2 datasets (GSE16538 and GSE20050), and 172 SARC-only DEGs in GSE16538 through Venn diagrams software. Different color meant different datasets. (A) 55 DEGs were upregulated in the 2 datasets, 142 DEGs were in GSE16238 (logFC >0). (B) 1 DEG was downregulated in 2 datasets, 30 DEGs in GSE16238 (logFC <0). Different color meant different datasets. DEGs – differentially expressed genes; SARC – sarcoidosis; PTB – pulmonary tuberculosis. 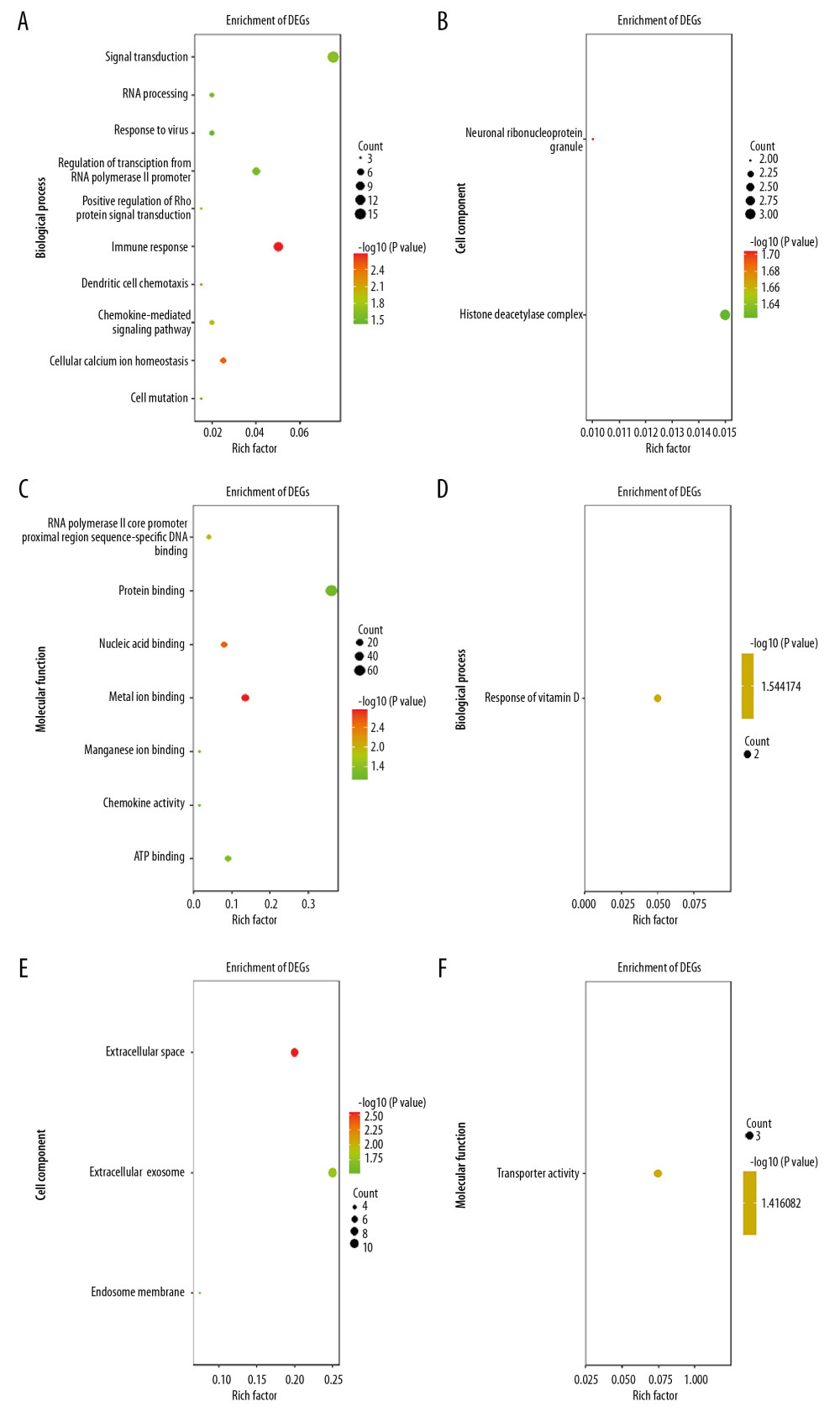 Figure 2. GO terms of overlapped SARC-only DEGs. (A–C) Upregulated genes. (D–F) Downregulated genes. (A, D), biological process; (B, E), cellular component; (C, F), molecular function. GO – Gene ontology; DEGs – differentially expressed genes; SARC – sarcoidosis.
Figure 2. GO terms of overlapped SARC-only DEGs. (A–C) Upregulated genes. (D–F) Downregulated genes. (A, D), biological process; (B, E), cellular component; (C, F), molecular function. GO – Gene ontology; DEGs – differentially expressed genes; SARC – sarcoidosis. 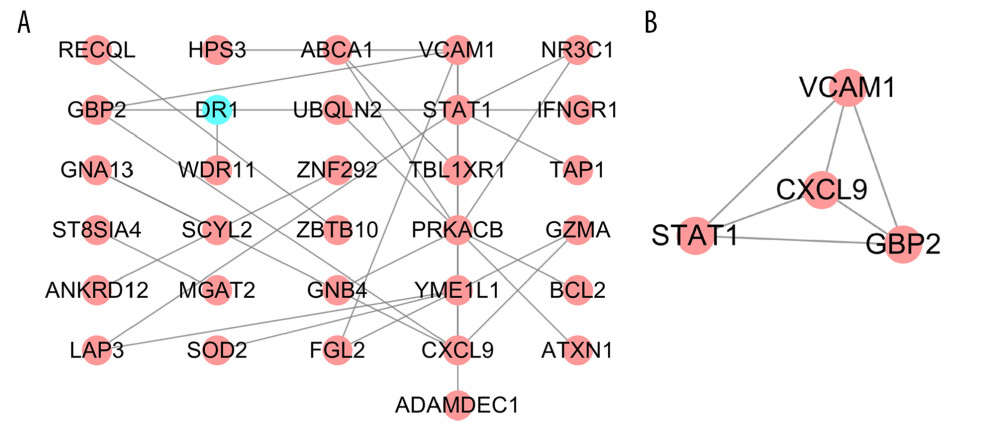 Figure 3. PPI network of SARC-PTB common DEGs and modules analysis. (A) PPI network of SARC-PTB common DEGs; (B) the only one module of PPI network. The red nodes represent the upregulated DEGs. The blue nodes represent the downregulated DEGs. The nodes meant proteins; the edges meant the interaction of proteins. PPI – protein–protein interaction; DEGs – differentially expressed genes; SARC – sarcoidosis; PTB – pulmonary tuberculosis.
Figure 3. PPI network of SARC-PTB common DEGs and modules analysis. (A) PPI network of SARC-PTB common DEGs; (B) the only one module of PPI network. The red nodes represent the upregulated DEGs. The blue nodes represent the downregulated DEGs. The nodes meant proteins; the edges meant the interaction of proteins. PPI – protein–protein interaction; DEGs – differentially expressed genes; SARC – sarcoidosis; PTB – pulmonary tuberculosis. 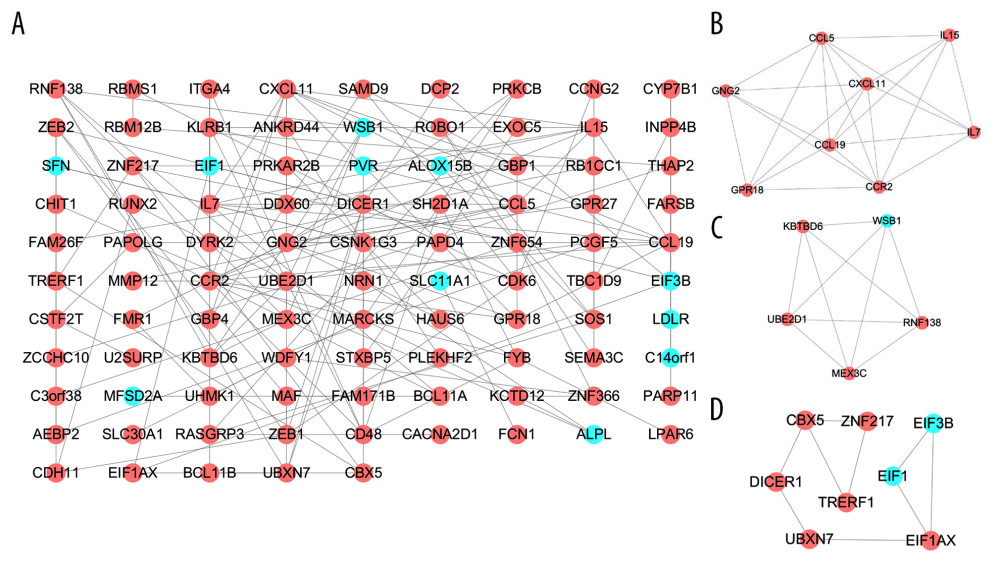 Figure 4. PPI network of SARC-only DEGs and modules analysis. (A) PPI network of SARC-only DEGs; (B–D) the top 3 modules of PPI network. The red nodes represent the upregulated DEGs. The blue nodes represent the downregulated DEGs. The nodes meant proteins; the edges meant the interaction of proteins. PPI – protein–protein interaction; DEG – differentially expressed genes; SARC – sarcoidosis.
Figure 4. PPI network of SARC-only DEGs and modules analysis. (A) PPI network of SARC-only DEGs; (B–D) the top 3 modules of PPI network. The red nodes represent the upregulated DEGs. The blue nodes represent the downregulated DEGs. The nodes meant proteins; the edges meant the interaction of proteins. PPI – protein–protein interaction; DEG – differentially expressed genes; SARC – sarcoidosis. 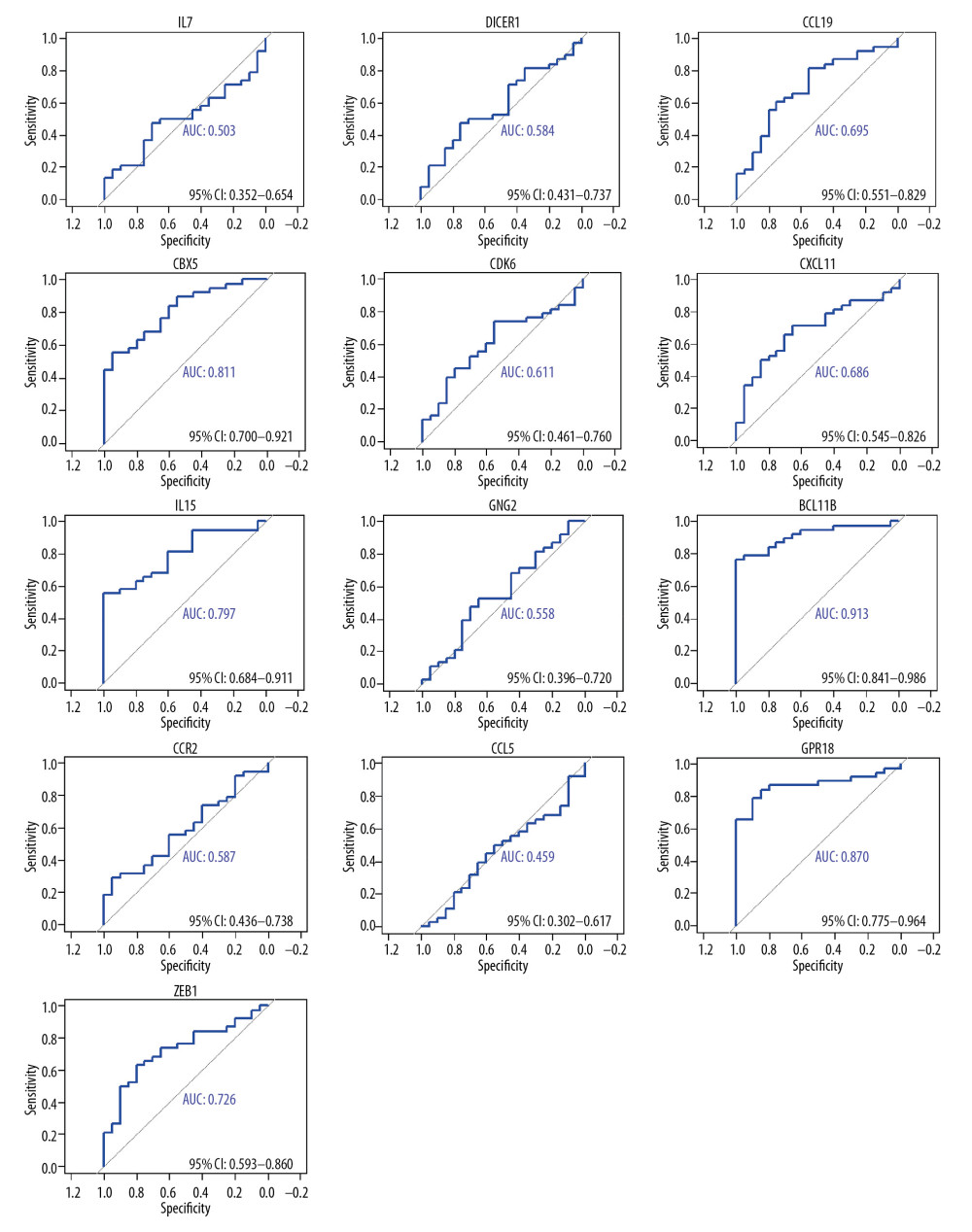 Figure 5. The diagnosis information of the 13 hub genes. The ROC curves were used to identify the diagnosis information of the 13 hub genes and, except CCL5. CBX5, BCL11B and GPR18 had a significantly diagnostic value (AUC >0.80). ROC curve – receiver operating characteristic curve; AUC – the area under the ROC curve.
Figure 5. The diagnosis information of the 13 hub genes. The ROC curves were used to identify the diagnosis information of the 13 hub genes and, except CCL5. CBX5, BCL11B and GPR18 had a significantly diagnostic value (AUC >0.80). ROC curve – receiver operating characteristic curve; AUC – the area under the ROC curve. Tables
Table 1. The detail information of 3 GEO datasets.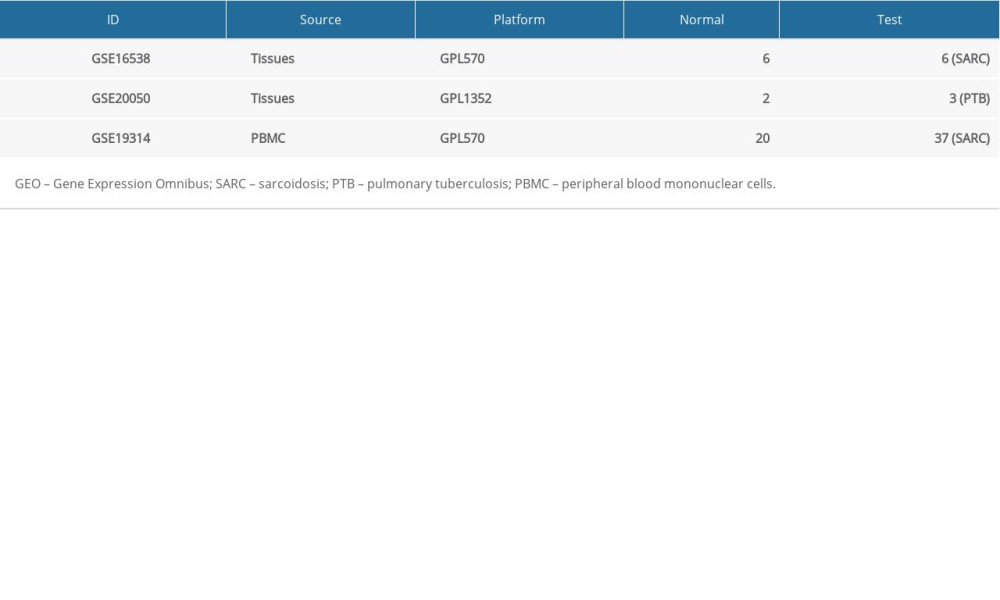 Table 2. SARC-PTB common DEGs.
Table 2. SARC-PTB common DEGs.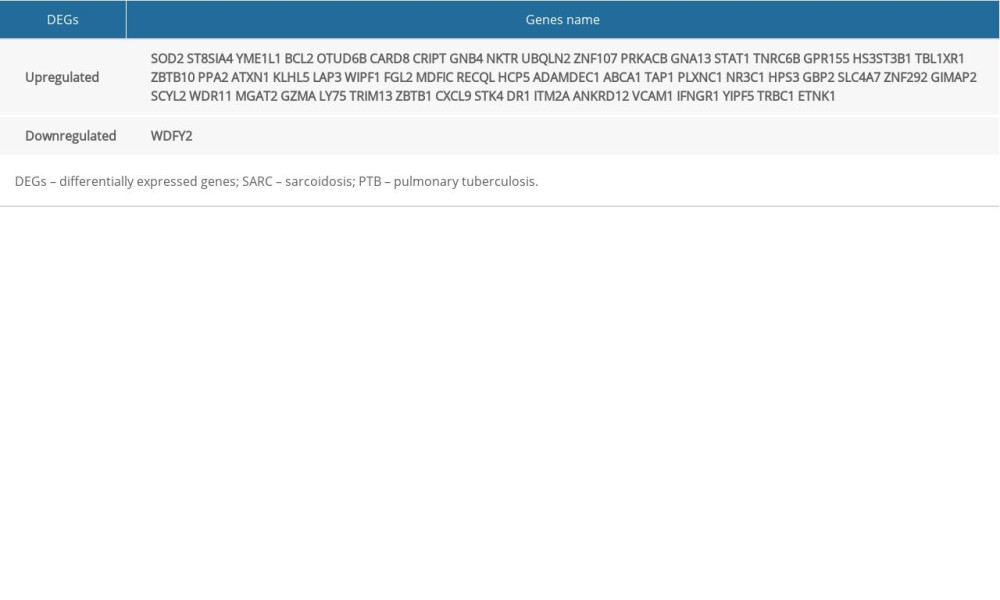 Table 3. SARC-only DEGs.
Table 3. SARC-only DEGs.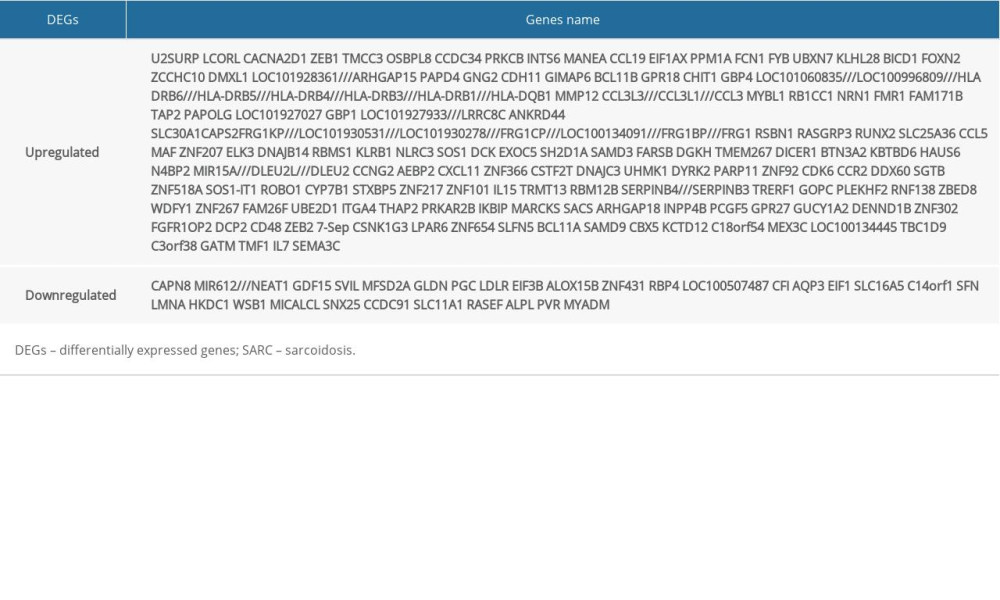 Table 4. GO analysis of SARC-PTB common DEGs.
Table 4. GO analysis of SARC-PTB common DEGs.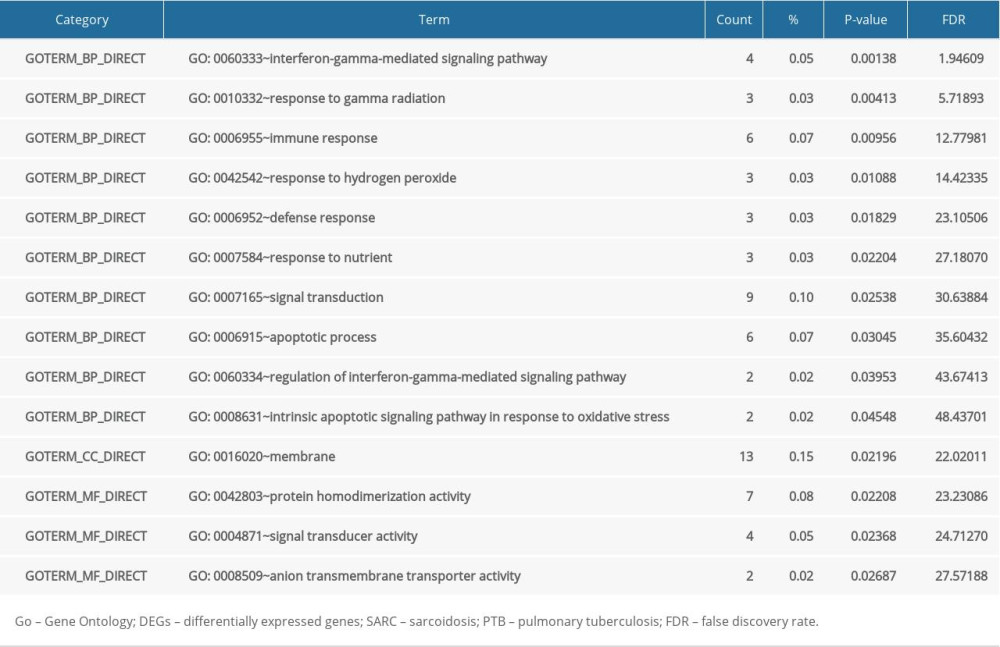 Table 5. KEGG pathway analysis of SARC-PTB common DEGs.
Table 5. KEGG pathway analysis of SARC-PTB common DEGs.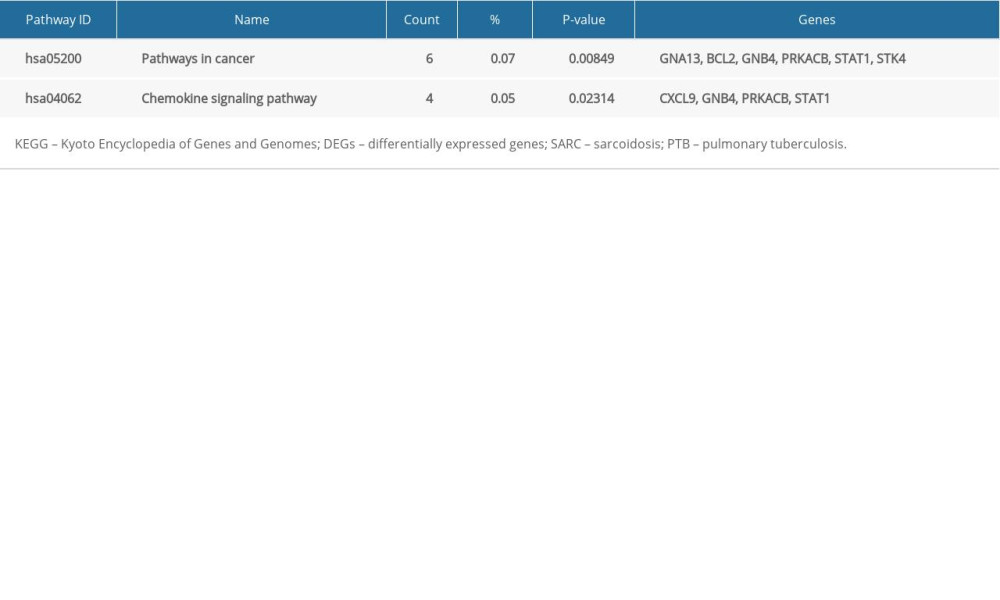 Table 6. KEGG pathway analysis of SARC-only DEGs.
Table 6. KEGG pathway analysis of SARC-only DEGs.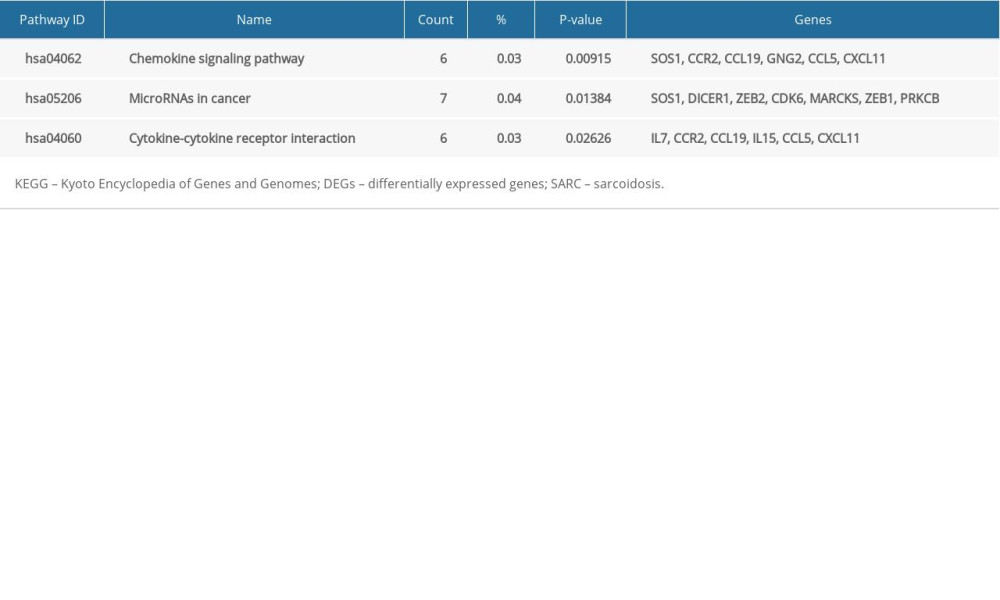 Table 7. The top 10 genes in each sub-plugin of SARC-only DEGs by cytoHubba.
Table 7. The top 10 genes in each sub-plugin of SARC-only DEGs by cytoHubba.
References
1. Landi C, Bargagli E, Carleo A, A functional proteomics approach to the comprehension of sarcoidosis: J Proteomics, 2015; 128; 375-87
2. Perez RL, Rivera-Marrero CA, Roman J, Pulmonary granulomatous inflammation: From sarcoidosis to tuberculosis: Semin Respir Infect, 2003; 18(1); 23-32
3. Gupta D, Agarwal R, Aggarwal AN, Jindal SK, Molecular evidence for the role of mycobacteria in sarcoidosis: A meta-analysis: Eur Respir J, 2007; 30(3); 508-16
4. Sharma OP, Murray Kornfeld, American College of Chest Physician, and sarcoidosis: A historical footnote: 2004 Murray Kornfeld Memorial Founders Lecture: Chest, 2005; 128(3); 1830-35
5. Li QH, Zhang Y, Zhao MM, Simultaneous amplification and testing method for Mycobacterium tuberculosis rRNA to differentiate sputum-negative tuberculosis from sarcoidosis: Am J Physiol Lung Cell Mol Physiol, 2019; 316(3); L519-24
6. Maertzdorf J, Weiner J, Mollenkopf HJ, Common patterns and disease-related signatures in tuberculosis and sarcoidosis: Proc Natl Acad Sci USA, 2012; 109(20); 7853-58
7. Guo T, Ma H, Zhou Y, Bioinformatics analysis of microarray data to identify the candidate biomarkers of lung adenocarcinoma: Peer J, 2019; 7; e7313
8. Landi C, Carleo A, Cillis G, Rottoli P, Sarcoidosis: Proteomics and new perspectives for improving personalized medicine: Expert Rev Proteomics, 2018; 15(10); 829-35
9. Kim MJ, Wainwright HC, Locketz M, Caseation of human tuberculosis granulomas correlates with elevated host lipid metabolism: Embo Mol Med, 2010; 2(7); 258-74
10. Katti MK, Assessment of serum IL-1, IL-2 and IFN-gamma levels in untreated pulmonary tuberculosis patients: Role in pathogenesis: Arch Med Res, 2011; 42(3); 199-201
11. Hong S, Mok Y, Jeon C, Tuberculosis, smoking and risk for lung cancer incidence and mortality: Int J Cancer, 2016; 139(11); 2447-55
12. Alberg AJ, Brock MV, Ford JG: Chest, 2013; 143(5 Suppl); e1S-e29S
13. O’Callaghan DS, O’Donnell D, O’Connell F, O’Byrne KJ, The role of inflammation in the pathogenesis of non-small cell lung cancer: J Thorac Oncol, 2010; 5(12); 2024-36
14. Chang SH, Mirabolfathinejad SG, Katta H, T helper 17 cells play a critical pathogenic role in lung cancer: Proc Natl Acad Sci USA, 2014; 111(15); 5664-69
15. Sato T, Arai E, Kohno T, Epigenetic clustering of lung adenocarcinomas based on DNA methylation profiles in adjacent lung tissue: Its correlation with smoking history and chronic obstructive pulmonary disease: Int J Cancer, 2014; 135(2); 319-34
16. Chaturvedi AK, Caporaso NE, Katki HA, C-reactive protein and risk of lung cancer: J Clin Oncol, 2010; 28(16); 2719-26
17. Kumar S, Baghdadi S, Cale AR, Concurrence of sarcoidosis and lung cancer: A diagnostic dilemma: Thorax, 2006; 61(12); 1100
18. McNeill M, Zanders TB, Morris MJ, A 49-year-old man with concurrent diagnoses of lung cancer, sarcoidosis, and multiple regions of adenopathy on positron emission tomography: Chest, 2009; 135(2); 546-49
19. Tokuyasu H, Izumi H, Mukai N, Small cell lung cancer complicated by pulmonary sarcoidosis: Intern Med, 2010; 49(18); 1997-2001
20. Kachalia AG, Ochieng P, Kachalia K, Rahman H, Rare coexistence of sarcoidosis and lung adenocarcinoma: Respir Med Case Rep, 2014; 12; 4-6
21. Iijima Y, Sugiyama Y, Sawahata M, Clinical features of pulmonary sarcoidosis complicated by lung cancer: Intern Med, 2017; 56(15); 1957-60
22. Tsuchiya K, Karayama M, Sato T, Simultaneous occurrence of sarcoidosis and anti-neutrophil cytoplasmic antibody-associated vasculitis in a patient with lung cancer: Intern Med, 2019; 58(22); 3299-304
23. Svendsen CB, Milman N, Nielsen HW, A prospective study evaluating the presence of Rickettsia in Danish patients with sarcoidosis: Scand J Infect Dis, 2009; 41(10); 745-52
24. Planck A, Eklund A, Grunewald J, Vene S: Eur Respir J, 2004; 24(5); 811-13
25. Biberfeld P, Petren AL, Eklund A, Human herpesvirus-6 (HHV-6, HBLV) in sarcoidosis and lymphoproliferative disorders: J Virol Methods, 1988; 21(1–4); 49-59
26. Moravvej H, Vesal P, Abolhasani E, Comorbidity of leishmania major with cutaneous sarcoidosis: Indian J Dermatol, 2014; 59(3); 316
27. Esteves T, Aparicio G, Garcia-Patos V, Is there any association between Sarcoidosis and infectious agents?: A systematic review and meta-analysis: BMC Pulm Med, 2016; 16(1); 165
28. Derler AM, Eisendle K, Baltaci M, High prevalence of ‘Borrelia-like’ organisms in skin biopsies of sarcoidosis patients from Western Austria: J Cutan Pathol, 2009; 36(12); 1262-68
29. Martens H, Zollner B, Zissel G: Eur Respir J, 1997; 10(6); 1356-58
30. Niimi T, Tomita H, Sato S, Vitamin D receptor gene polymorphism in patients with sarcoidosis: Am J Respir Crit Care Med, 1999; 160(4); 1107-9
31. Stjepanovic MI, Mihailovic-Vucinic V, Spasovski V, Genes and metabolic pathway of sarcoidosis: Identification of key players and risk modifiers: Arch Med Sci, 2019; 15(5); 1138-46
32. Riddle EL, Schwartzman RA, Bond M, Insel PA, Multi-tasking RGS proteins in the heart: the next therapeutic target?: Circ Res, 2005; 96(4); 401-11
33. van Dijk N, Sprangers MA, Boer KR, Quality of life within one year following presentation after transient loss of consciousness: Am J Cardiol, 2007; 100(4); 672-76
34. Huang YJ, Zhou ZW, Xu M, Alteration of gene expression profiling including GPR174 and GNG2 is associated with vasovagal syncope: Pediatr Cardiol, 2015; 36(3); 475-80
35. Gibejova A, Mrazek F, Subrtova D, Expression of macrophage inflammatory protein-3 beta/CCL19 in pulmonary sarcoidosis: Am J Respir Crit Care Med, 2003; 167(12); 1695-703
36. Kriegova E, Tsyrulnyk A, Arakelyan A, Expression of CCX CKR in pulmonary sarcoidosis: Inflamm Res, 2006; 55(10); 441-45
37. Arger NK, Ho M, Woodruff PG, Koth LL, Serum CXCL11 correlates with pulmonary outcomes and disease burden in sarcoidosis: Respir Med, 2019; 152; 89-96
38. Guo J, Zhang Z, Pan L, Zhou Y, Identification of miR-758-3p as potential modulator of CBX5 expression in gastric cancer: Technol Cancer Res Treat, 2018; 17 1533033818816061
39. Wang J, Yang Z, Wu NP, Yang J, Increased expression of BCL11B and its recruited chromatin remodeling factors during highly active antiretroviral therapy synergistically represses the transcription of human immunodeficiency virus type 1 and is associated with residual immune activation: Arch Virol, 2020; 165(2); 321-30
40. Nourbakhsh F, Atabaki R, Roohbakhsh A, The role of orphan G protein-coupled receptors in the modulation of pain: A review: Life Sci, 2018; 212; 59-69
Figures
 Figure 1. Identification of 56 common SARC-PTB DEGs in the 2 datasets (GSE16538 and GSE20050), and 172 SARC-only DEGs in GSE16538 through Venn diagrams software. Different color meant different datasets. (A) 55 DEGs were upregulated in the 2 datasets, 142 DEGs were in GSE16238 (logFC >0). (B) 1 DEG was downregulated in 2 datasets, 30 DEGs in GSE16238 (logFC <0). Different color meant different datasets. DEGs – differentially expressed genes; SARC – sarcoidosis; PTB – pulmonary tuberculosis.
Figure 1. Identification of 56 common SARC-PTB DEGs in the 2 datasets (GSE16538 and GSE20050), and 172 SARC-only DEGs in GSE16538 through Venn diagrams software. Different color meant different datasets. (A) 55 DEGs were upregulated in the 2 datasets, 142 DEGs were in GSE16238 (logFC >0). (B) 1 DEG was downregulated in 2 datasets, 30 DEGs in GSE16238 (logFC <0). Different color meant different datasets. DEGs – differentially expressed genes; SARC – sarcoidosis; PTB – pulmonary tuberculosis. Figure 2. GO terms of overlapped SARC-only DEGs. (A–C) Upregulated genes. (D–F) Downregulated genes. (A, D), biological process; (B, E), cellular component; (C, F), molecular function. GO – Gene ontology; DEGs – differentially expressed genes; SARC – sarcoidosis.
Figure 2. GO terms of overlapped SARC-only DEGs. (A–C) Upregulated genes. (D–F) Downregulated genes. (A, D), biological process; (B, E), cellular component; (C, F), molecular function. GO – Gene ontology; DEGs – differentially expressed genes; SARC – sarcoidosis. Figure 3. PPI network of SARC-PTB common DEGs and modules analysis. (A) PPI network of SARC-PTB common DEGs; (B) the only one module of PPI network. The red nodes represent the upregulated DEGs. The blue nodes represent the downregulated DEGs. The nodes meant proteins; the edges meant the interaction of proteins. PPI – protein–protein interaction; DEGs – differentially expressed genes; SARC – sarcoidosis; PTB – pulmonary tuberculosis.
Figure 3. PPI network of SARC-PTB common DEGs and modules analysis. (A) PPI network of SARC-PTB common DEGs; (B) the only one module of PPI network. The red nodes represent the upregulated DEGs. The blue nodes represent the downregulated DEGs. The nodes meant proteins; the edges meant the interaction of proteins. PPI – protein–protein interaction; DEGs – differentially expressed genes; SARC – sarcoidosis; PTB – pulmonary tuberculosis. Figure 4. PPI network of SARC-only DEGs and modules analysis. (A) PPI network of SARC-only DEGs; (B–D) the top 3 modules of PPI network. The red nodes represent the upregulated DEGs. The blue nodes represent the downregulated DEGs. The nodes meant proteins; the edges meant the interaction of proteins. PPI – protein–protein interaction; DEG – differentially expressed genes; SARC – sarcoidosis.
Figure 4. PPI network of SARC-only DEGs and modules analysis. (A) PPI network of SARC-only DEGs; (B–D) the top 3 modules of PPI network. The red nodes represent the upregulated DEGs. The blue nodes represent the downregulated DEGs. The nodes meant proteins; the edges meant the interaction of proteins. PPI – protein–protein interaction; DEG – differentially expressed genes; SARC – sarcoidosis. Figure 5. The diagnosis information of the 13 hub genes. The ROC curves were used to identify the diagnosis information of the 13 hub genes and, except CCL5. CBX5, BCL11B and GPR18 had a significantly diagnostic value (AUC >0.80). ROC curve – receiver operating characteristic curve; AUC – the area under the ROC curve.
Figure 5. The diagnosis information of the 13 hub genes. The ROC curves were used to identify the diagnosis information of the 13 hub genes and, except CCL5. CBX5, BCL11B and GPR18 had a significantly diagnostic value (AUC >0.80). ROC curve – receiver operating characteristic curve; AUC – the area under the ROC curve. Tables
 Table 1. The detail information of 3 GEO datasets.
Table 1. The detail information of 3 GEO datasets. Table 2. SARC-PTB common DEGs.
Table 2. SARC-PTB common DEGs. Table 3. SARC-only DEGs.
Table 3. SARC-only DEGs. Table 4. GO analysis of SARC-PTB common DEGs.
Table 4. GO analysis of SARC-PTB common DEGs. Table 5. KEGG pathway analysis of SARC-PTB common DEGs.
Table 5. KEGG pathway analysis of SARC-PTB common DEGs. Table 6. KEGG pathway analysis of SARC-only DEGs.
Table 6. KEGG pathway analysis of SARC-only DEGs. Table 7. The top 10 genes in each sub-plugin of SARC-only DEGs by cytoHubba.
Table 7. The top 10 genes in each sub-plugin of SARC-only DEGs by cytoHubba. Table 1. The detail information of 3 GEO datasets.
Table 1. The detail information of 3 GEO datasets. Table 2. SARC-PTB common DEGs.
Table 2. SARC-PTB common DEGs. Table 3. SARC-only DEGs.
Table 3. SARC-only DEGs. Table 4. GO analysis of SARC-PTB common DEGs.
Table 4. GO analysis of SARC-PTB common DEGs. Table 5. KEGG pathway analysis of SARC-PTB common DEGs.
Table 5. KEGG pathway analysis of SARC-PTB common DEGs. Table 6. KEGG pathway analysis of SARC-only DEGs.
Table 6. KEGG pathway analysis of SARC-only DEGs. Table 7. The top 10 genes in each sub-plugin of SARC-only DEGs by cytoHubba.
Table 7. The top 10 genes in each sub-plugin of SARC-only DEGs by cytoHubba. In Press
07 Mar 2024 : Clinical Research
Knowledge of and Attitudes Toward Clinical Trials: A Questionnaire-Based Study of 179 Male Third- and Fourt...Med Sci Monit In Press; DOI: 10.12659/MSM.943468
08 Mar 2024 : Animal Research
Modification of Experimental Model of Necrotizing Enterocolitis (NEC) in Rat Pups by Single Exposure to Hyp...Med Sci Monit In Press; DOI: 10.12659/MSM.943443
18 Apr 2024 : Clinical Research
Comparative Analysis of Open and Closed Sphincterotomy for the Treatment of Chronic Anal Fissure: Safety an...Med Sci Monit In Press; DOI: 10.12659/MSM.944127
08 Mar 2024 : Laboratory Research
Evaluation of Retentive Strength of 50 Endodontically-Treated Single-Rooted Mandibular Second Premolars Res...Med Sci Monit In Press; DOI: 10.12659/MSM.944110
Most Viewed Current Articles
17 Jan 2024 : Review article
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
14 Dec 2022 : Clinical Research
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387
01 Jan 2022 : Editorial
Editorial: Current Status of Oral Antiviral Drug Treatments for SARS-CoV-2 Infection in Non-Hospitalized Pa...DOI :10.12659/MSM.935952
Med Sci Monit 2022; 28:e935952








