24 May 2021: Meta-Analysis
Relationship of Programmed Death-1 (PD-1) and Programmed Death Ligand-1 (PD-L1) Polymorphisms with Overall Cancer Susceptibility: An Updated Meta-Analysis of 28 Studies with 60 612 Subjects
Wenjing Zhang12BCDEF, Yuxuan Song34ABEF, Xiangcheng Zhang1ACEG*DOI: 10.12659/MSM.932146
Med Sci Monit 2021; 27:e932146
Abstract
BACKGROUND: Programmed death-1 and its ligand-1 (PD-1/PD-L1) regulate tumor immunotherapy. A large number of studies have explored the relationship between PD-1, PD-L1, and different tumor susceptibility. However, these conclusions are not always consistent. Therefore, we updated this meta-analysis.
MATERIAL AND METHODS: MEDLINE, Web of Science, EMBASE and other databases were searched systematically to obtain related research. Then, we used STATA15.0 software to carry out the final meta-analysis. The computational advantage is better than OR to evaluate this relationship.
RESULTS: A total of a total of 28 related studies were involved in our meta-analysis. It was found that PD-1 rs11568821 and rs7421861 increased the overall cancer probability in the allelic genetic model, while PD-1 rs36084323 effectively reduced the risk of cancer in the dominant genetic model. In the homozygous genetic model, PD-L1 rs17718883 effectively increased the probability of tumorigenesis. PD-L1rs4143815 is associated with a reduced incidence of cancer in heterozygote, homozygote and dominant genetic patterns. Subgroup analysis showed that PD-1rs2227981 can promote the susceptibility to breast cancer, while PD-1rs2227982 can reduce the susceptibility to breast cancer. PD-L1 rs2890658 can significantly reduce the risk of lung and liver cancer.
CONCLUSIONS: PD-1rs11568821, rs36084323, rs7421861, pD-L1rs17718883, and rs4143815 are associated with tumor susceptibility. However, a review based on more experimental evidence is needed to verify our findings.
Keywords: Disease Susceptibility, Genes, Neoplasm, PDCD1 Protein, Human, Alleles, B7-H1 Antigen, Genetic Predisposition to Disease, Immunotherapy, Incidence, Neoplasms, Polymorphism, Single Nucleotide, Programmed Cell Death 1 Receptor
Background
Cancer is a serious global problem. According to the American Cancer Society (ACS), there were more than 1.8 million new cancer cases and 606 520 cancer-related deaths in the United States last year [1]. In the past few decades, considerable progress has been made in understanding how cancer successfully escapes from the immune system and survives, and these advances provide an active way to overcome tumor immune escape and may help to eliminate cancer cells [2]. As a preliminary study, immunotherapy is mainly concentrated in immune checkpoints [3]. Programmed cell death protein 1 and its ligand (PD-1/PD-L1) pathway play an important role in the induction and maintenance of immune tolerance in the tumor microenvironment [4]. Early studies have shown that several IgG4 antibodies point to PD-1/PD-L1 in some solid tumors. These studies have helped to improve the first round of PD-1 inhibitors, such as nivolumab, which were approved by the U.S. Food and Drug Administration (FDA) in 2014 [5]. PD-L1, pointing to PD-1, on T cells helps to destroy cancer cells. However, tumor cells show immune escape through the expression of PD-L1 [6]. The overexpression of PD-L1 in many kinds of cells, such as cancer cells and antigen-presenting cell (APC), is considered to be the key factor for maintaining anti-tumor immunity in the tumor microenvironment (TME) and tumor draining lymph nodes. The increase of PD-L1 level is related to the enhancement of (ICB) response blocked by immune checkpoints against PD-1/PD-L1 [7]. Due to the complexity of tumor immunity, there is no definite biomarker to evaluate the results of PD-1/PD-L1 targeted therapy [8]. It is very important to identify the single-nucleotide polymorphism (SNP) that affects expression of the PD-1 gene and participates in tumor susceptibility. Therefore, it will help to predict potential individuals and clarify the pathophysiological mechanism of cancer [9]. A growing number of studies have explored the relationship between PD-1 and PD-L1 single-nucleotide polymorphisms and multiple cancer susceptibility. However, their results are not consistent, and the same location has different effects in different studies [10–19]. More systematic reviews are needed to support the evidence of these results. To address this problem, we propose a new meta-analysis to assess the relationship between PD-1 and PD-L1 and cancer susceptibility.
Material and Methods
LITERATURE SEARCH:
The related literatures in the databases of PubMed, Web of Science, EMBASE, and China National knowledge Infrastructure (CNKI) and Wanfang data information Service platform were searched by computer, and the related research was carried out. To determine the relationship between PD-1/PD-L1 mutations and cancer susceptibility, we used the following keywords: (programmed cell death 1 or programmed cell death ligand1 or PD-1 or PDCD1 or PD-L1 or CD274 or B7-H1) and (polymorphism or genotype or variant or SNP) and (cancer or carcinoma or neoplasm).
We searched for relevant published research until March 28, 2019, and we also searched the literature for relevant dissertations. Figure 1 shows a flowchart of search strategies that illustrate PD-1 and PD-L1 variants and cancer predisposition.
INCLUSION AND EXCLUSION CRITERIA:
The process of retrieving studies is shown in Figure 1. We list the inclusion and exclusion standards below.
INCLUSION CRITERIA:
The inclusion criteria were: (1) Case-control studies on the relationship of PD-1 and PD-L1 variant with cancer predisposition; (2) The genotypes of control groups were in Hardy-Weinberg equilibrium (HWE); (3) Allele frequencies in studies were shown; (4) English and Chinese articles; and (5) Studies with human subjects.
EXCLUSION CRITERIA:
The exclusion criteria were as follows: (1) genotypes in the control group did not conform to HWE; (2) studies of genotype frequency estimates of odds ratio (OR) and 95% confidence interval (CI) could not be obtained; (3) useful data or results could not be extracted; (4) the results were not related to cancer susceptibility; (5) the article was a duplicate publication or existed only as an abstract.
DATA EXTRACTION:
The data were extracted independently by 2 authors. If necessary, the differences between the 2 authors were resolved through discussion with a third investigator. All valid data are shown in Table 1: author’s name, year, country, nationality, cancer type, number of groups, P value of HWE, and genotype method.
TRIALS QUALITY ASSESSMENT:
The quality of the selected articles was evaluated with the Ottawa Newcastle scale (NOS). The NOS score was ranged from 0 to 9. Scores greater than 5 indicate high-quality articles. Table 2 lists the quality of all selected studies. The assessment includes the following 3 parts: (1) selection of subjects; (2) comparability between groups; and (3) exposure assessment.
STATISTICAL ANALYSES:
STATA15.0 software was used for statistical analysis. Q test and heterogeneity coefficient I2 were used to evaluate the heterogeneity of the study. If there was no statistical heterogeneity (I2 <50%), our meta-analysis used a fixed-effect model, as well as random effects. The combined OR value and its 95% CI were evaluated to assess the relationship. We include 5 genetic models: (1) allele, (2) heterozygote, (3) homozygote, (4) dominance, and (5) recessive model, and studies the relationship between them. If it was indispensable, we analyzed the relevant causes that may have led to heterogeneity in groups. We used the funnel chart and the P value of Egger’s and Begg’s test to judge the publication deviation.
Results
CHARACTERISTICS OF THE SELECTED STUDIES:
The characteristics of the selected articles [9,20–41] are shown in Table 1. Twenty-eight articles were identified for the relationship between PD-1/PD-L1 SNP and cancers susceptibility. These studies contained 60 612 subjects.
A total of 25 articles were involved in the meta-analysis of PD-1 SNPs. A total of 6 trials were conducted to study the relationship between PD-1 rs10204525 and tumor susceptibility, with 8940 subjects. Five studies involving 3156 subjects investigated the relationship between PD-1 rs11568821 and cancer predisposition. A total of 7829 subjects in 11 studies showed a relationship between PD-1rs2227981 mutations and cancer susceptibility. Ten trials, including 10 976 subjects, investigated the relationship between PD-1rs2227982 and cancer susceptibility. A total of 11 005 subjects in 10 studies reported the relationship between PD-1 rs36084323 and cancer susceptibility. A total of 10 963 subjects in 8 studies analyzed the relationship between PD-1 rs7421861 variation and cancer susceptibility. For PD-L1 SNPs, a total of 8 articles that studied the effects of 3 widely studied polymorphisms in PD-L1 were included in the meta-analysis. Three studies were conducted to assess the relationship between PD-L1 rs17718883 mutations and cancer susceptibility. Six trials involving 3797 subjects confirmed the relationship between PD-L1 rs2890658 and cancer susceptibility. Six studies, including 3015 subjects, estimated the relationship between PD-L1 rs4143815 and cancer susceptibility.
META-ANALYSIS RESULTS:
Table 3 contains our meta-analysis summary of PD-1 and PD-L1 variants and cancer susceptibility. Table 4 lists the subgroup analyses based on cancer type.
ORS OF PD-1 SNPS AND CANCER PREDISPOSITION: As regards PD-1 rs11568821 polymorphism, we revealed the variant increased the cancer predisposition in the allele genetic model (OR=1.314, 95% CI=1.116–1.547, P=0.001, G vs A). PD-1 rs36084323 variant was proved to decrease the cancer risk in dominant genetic model (OR=0.903, 95% CI =0.819–0.995, P=0.038, GG+GA vs AA). PD-1 rs7421861 variant was found to enhance cancer predisposition in the heterozygote model (OR=1.202, 95% CI=1.031–1.402, P=0.019, TC vs CC) and dominant genetic model (OR=1.181, 95% CI=1.020–1.368, P=0.026, TT+CT vs CC). No clear relationship was found between rs2227981, rs227982, and rs10204525 variants and cancer susceptibility. Forest plots of meta-analysis of PD-1 rs11568821 and rs10204525 in the allele model are demonstrated in Figure 2. Forest plots of meta-analysis on PD-1 rs36084323 and rs2227981 in the dominant model are shown in Figure 3. Forest plots of the meta-analysis of PD-1 rs7421861 in the heterozygote model and dominant model are presented in Figure 4.
SUBGROUP ANALYSIS OF PD-1 SNPS AND THE CANCER PREDISPOSITION: We conducted some subgroup analyses that were based on cancer types. We detected PD-1 rs2227981 promoted the predisposition of breast cancer (OR=1.219, 95% CI=1.045–1.422, p=0.012, C vs T, Figure 5). PD-1 rs2227982 variant was confirmed to decrease breast cancer risk in the allele model (OR=1.173, 95% CI=1.040–1.322, P=0.010, C vs T, Figure 5); homozygote model (OR=1.379, 95% CI=1.081–1.758, P=0.010, CC vs TT) and recessive genetic model (OR=1.375, 95% CI=1.126–1.679, P=0.002, CC vs CT+TT). As regards the PD-1 rs36084323, our analyses results showed this polymorphism lowered the ovarian cancer predisposition in the heterozygote model (OR=0.695, 95% CI=0.538–0.897, P=0.005, AG vs AA); homozygote model (OR=0.615, 95% CI=0.459–0.823, P=0.001, GG vs AA, Figure 6) and dominant genetic model (OR=0.666, 95% CI=0.523–0.847, P=0.001, GG+GA vs AA, Figure 6).
ORS OF PD-L1 SNPS AND CANCER PREDISPOSITION: As regards PD-L1 rs17718883 polymorphism, we demonstrated the variant increased the cancer predisposition in the homozygote genetic models (OR=25.488, 95% CI=8.494–76.481, P= 0, CC vs GG). PD-L1 rs4143815 was found to decrease cancer risk in the heterozygote genetic model (OR=0.560, 95% CI=0.365–0.860, P=0.008, GC vs GG), homozygote genetic model (OR=0.537, 95% CI= 0.315–0.918, P=0.023, CC vs GG), and dominant genetic model (OR=0.555, 95% CI= 0.351–0.877, P=0.012, CC+CG vs GG). There was no significant association of PD-L1 rs2890658 variant with cancer predisposition. Forest plots of meta-analysis on PD-L1 rs17718883 and rs2890658 in homozygote model are shown in Figure 7. Forest plots of meta-analysis about PD-L1 rs4143815 in heterozygote model and homozygote model are presented in Figure 8.
SUBGROUP ANALYSES OF PD-L1 SNPS AND THE CANCER PREDISPOSITION: We found that PD-L1 rs2890658 reduced the non-small cell lung cancer predisposition in the allele model (OR=0.609, 95% CI=0.477–0.777, P=0, A vs C) and recessive genetic model (OR=0.589, 95% CI=0.454–0.765, P=0, AA vs AC+CC). Moreover, PD-L1 rs2890658 variant increased the hepatocellular carcinoma predisposition in the allele model (OR=1.358, 95% CI=1.006–1.833, P=0.046, A vs C, Figure 9) and recessive genetic model (OR=1.405, 95% CI=1.002–1.970, P=0.049, AA vs AC+CC, Figure 9). With respect to PD-L1 rs4143815, we found that this polymorphism lowered the hepatocellular cancer predisposition in the heterozygote model (OR =0.422, 95% CI=0.279–0.639, P=0, GC vs GG, Figure 10), homozygote model (OR=0.459, 95% CI=0.213–0.991, p=0.047, CC vs GG, Figure 10), and dominant genetic model (OR=0.425, 95% CI=0.288–0.628, p=0, CC+CG vs GG).
SENSITIVITY ANALYSES AND PUBLICATION BIAS:
We performed a series of sensitivity analyses, and the composite results showed no significant change. This shows that our research results are reliable.
We used the P values of 2 tests to evaluate publication bias. Then, the results of merger analysis were compared in the allelic genetic model. The consequences of publication bias are listed in Table 5. If the P value of both tests is less than 0.05, it indicates that there is publication bias. In conclusion, there is no publication bias in the literature on the relationship between PD-1rs11568821, rs36094323, pD-L1 rs4143815, and rs7421861 gene polymorphisms and cancer risk. Figures 11 and 12 show Begg’s funnel plots of PD-1 rs11568821, rs36094323, rs7421861, and PD-L1 rs4143815 in the allele model, but there was publication bias about the relationship of PD-1 rs17718883 polymorphism and cancer predisposition in the allele genetic model.
Discussion
It has recently been confirmed that checkpoint blockade immunotherapy is one of the reasons for the continued decline in cancer mortality [1]. Single-nucleotide polymorphisms are expected to become biomarkers to help scientists classify tumors and will allow patients to be assigned to the most appropriate treatment [42]. PD-1 and PD-L1 play an indispensable role in immune tolerance, and they have become key targets in cancer therapy [43]. Some articles have discussed the relationship between PD-1 and PD-L1 mutations and different cancer susceptibility, but the conclusions are still inconsistent. Our study estimated the relationship between 9 variants of PD-1 and PD-L1 and cancer susceptibility. Our results suggest that PD-1 rs36084323 and PD-L1 rs4143815 are significantly associated with decreased cancer susceptibility, while PD-1 rs7421861, rs11568821, and PD-L1 rs17718883 variants increase overall cancer susceptibility. We found that there was no clear relationship between PD-1 rs2227981, rs2227982, rs10204525, and PD-L1rs2890658 mutations and cancer susceptibility.
Subgroup analysis showed that PD-1rs2227981 increased the susceptibility to breast cancer. In addition, PD-1rs2227982 is associated with reduced susceptibility to breast cancer. However, PD-1rs36084323 is associated with reduced susceptibility to ovarian cancer. In addition, our results suggest that the PD-L1rs4143815 mutation significantly reduces the susceptibility to liver cancer. There was a negative correlation between PD-L1rs2890658 and susceptibility to non-small cell lung cancer.
Recently, Hashemi et al [44] conducted a meta-analysis of 27 case-control studies to explore the relationship between PD-1 genes rs11568821, rs2227981, rs2227982, and rs7421861 and tumor susceptibility. The results showed that the mutations of PD-1rs2227981, rs11568821, rs7421861, and PD-L1rs4143815 were related to the overall susceptibility to cancer. However, Hashemi et al did not include Chinese studies, nor did they rule out low-quality studies based on NOS scores. Shan et al [10] conducted a meta-analysis of 10 studies (9571 subjects) and found that PD-1rs36084323 was associated with reduced susceptibility to cancer in Asians, which is consistent with our findings. Compared with the meta-analysis of Shan et al, the genetic polymorphism studied in this paper is more comprehensive and includes more recent research results. However, a study by Zhou et al [14] showed that PD-L1rs4143815 is associated with increased susceptibility to cancer, which is different from our findings. This phenomenon may be because Zhou Ju and others did not evaluate the quality of NOS studies or analyze the experiments that may be biased. Last but not least, Dong Wenjing et al [12] conducted a meta-analysis of 12 trials and confirmed that PD-1rs2227981 variation was associated with a significant decrease in cancer susceptibility. Compared with Dong Wenjing and others, the research included in our meta-analysis is newer and the scope of genetic polymorphism is wider. In addition, we suspected that the differences in the results of the study should be related to fa tors such as inclusion criteria and race.
Tumor immunotherapy targeting PD-1 and PD-L1 immune checkpoint pathways has begun in the field of oncology. The combination of anti-PD-1 and anti-cytotoxic T lymphocyte-associated protein 4 (CTLA-4) has produced an effective pathological response rate in the treatment of breast and lung cancer [45]. At present, 3 PD-L1 inhibitors have been approved for non-small cell lung cancer [46]. Aaron et al confirmed that the effective rate of blocking PD-1 with nivolumab was more than half in unselected patients with Hodgkin’s lymphoma [47]. Melanoma patients with other types of cancer showed the best response [48]. Our meta-analysis includes more case and control samples than previous studies, and we also included Chinese studies. In addition, we evaluated the quality of NOS research, including high-quality research and excluding low-quality articles. Last but not least, in our study, 26 of the 27 trials were conducted in Asians and only 1 in Whites, which could reduce the potential effects of different races on genetic susceptibility. Therefore, our meta-analysis makes a more convincing assessment than previous studies.
However, this study also has some limitations. First of all, the small number of participants with the PD-L1rs1771883 polymorphism may have led to a lack of statistical ability to study this relationship. Secondly, there is obvious heterogeneity in several polymorphisms; therefore, we conducted a subgroup analysis to find out the causes of heterogeneity. Finally, we inferred that the type of cancer and the country of residence of the participants may lead to heterogeneity. These mean that our results should be interpreted carefully.
Conclusions
In conclusion, our results suggest that both PD-1 rs36084323 and PD-L1 rs4143815 variants decrease cancer predisposition, while PD-1 rs7421861, rs11568821, and PD-L1 rs17718883 polymorphisms significantly increase the risk of cancer.
Figures
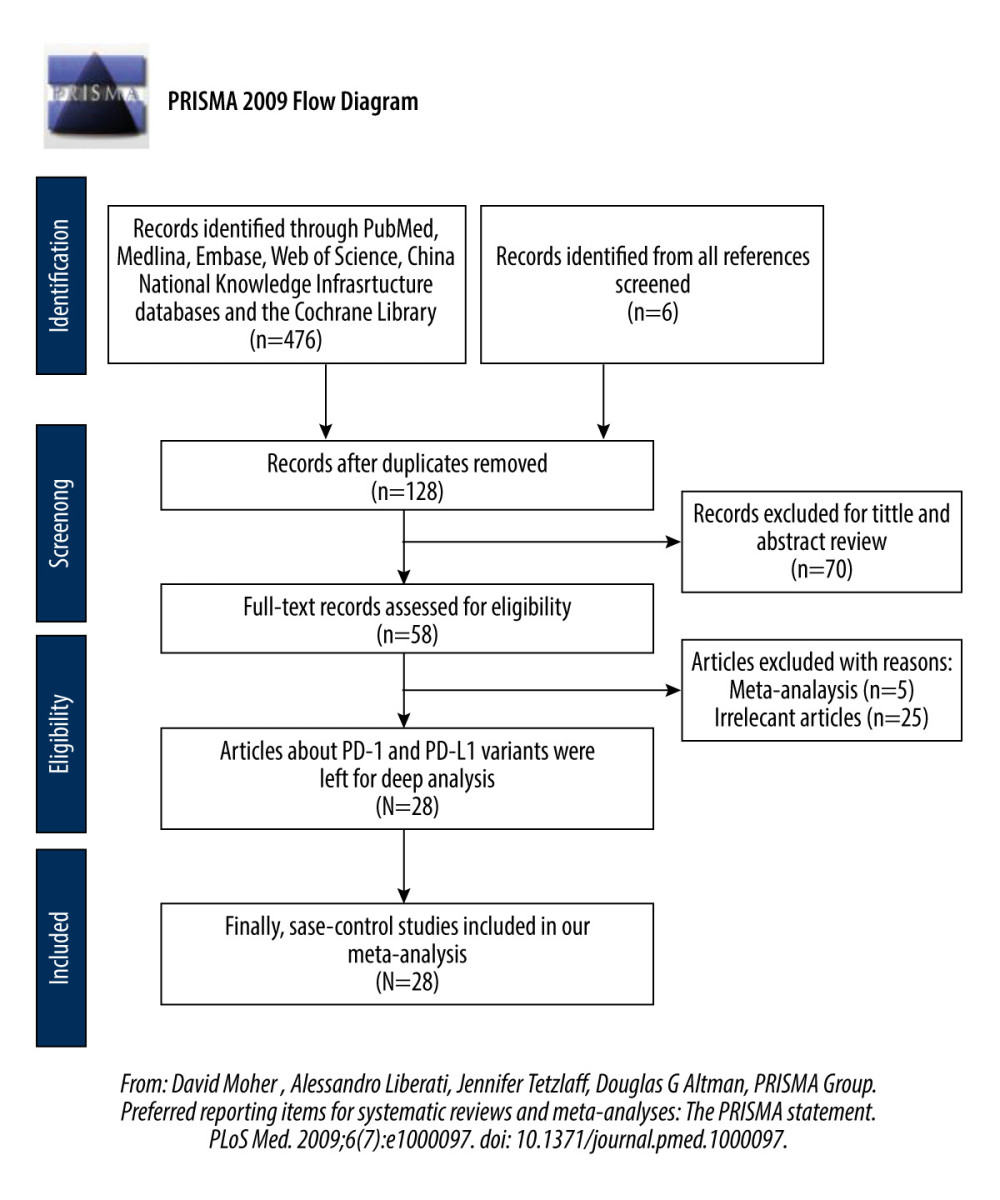 Figure 1. Flowchart illustrating the search strategy for PD-1 and PD-L1 variants and cancer.
Figure 1. Flowchart illustrating the search strategy for PD-1 and PD-L1 variants and cancer. 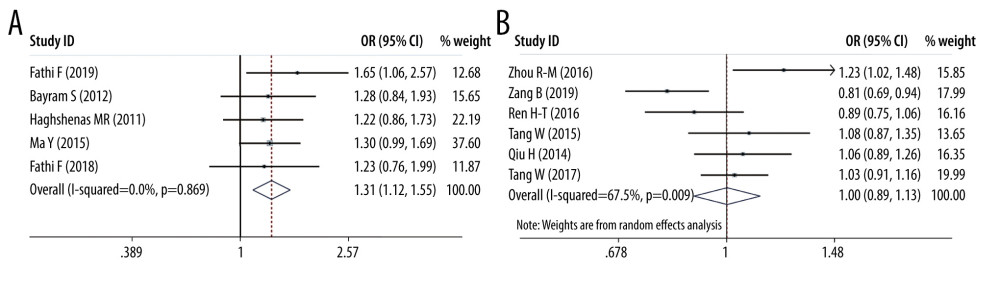 Figure 2. Forest plots of meta-analysis. (A) PD-1 rs11568821 in allele model (B) PD-1 rs10204525 in allele model.
Figure 2. Forest plots of meta-analysis. (A) PD-1 rs11568821 in allele model (B) PD-1 rs10204525 in allele model. 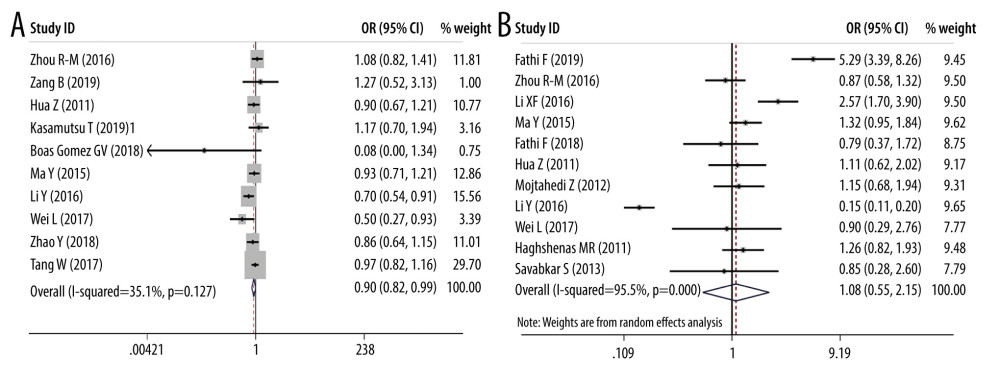 Figure 3. Forest plots of meta-analysis. (A) PD-1 rs36084323 in dominant model (B) PD-1 rs2227981 in dominant model.
Figure 3. Forest plots of meta-analysis. (A) PD-1 rs36084323 in dominant model (B) PD-1 rs2227981 in dominant model. 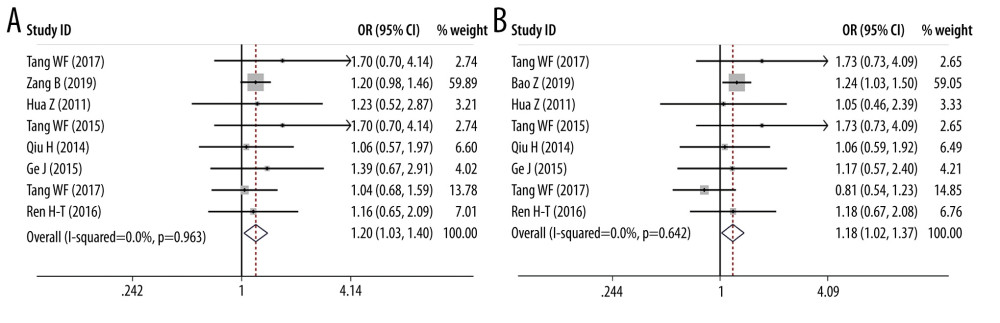 Figure 4. Forest plots of meta-analysis. (A) PD-1 rs7421861 in heterozygote model (B) PD-1 rs7421861 in dominant model.
Figure 4. Forest plots of meta-analysis. (A) PD-1 rs7421861 in heterozygote model (B) PD-1 rs7421861 in dominant model. 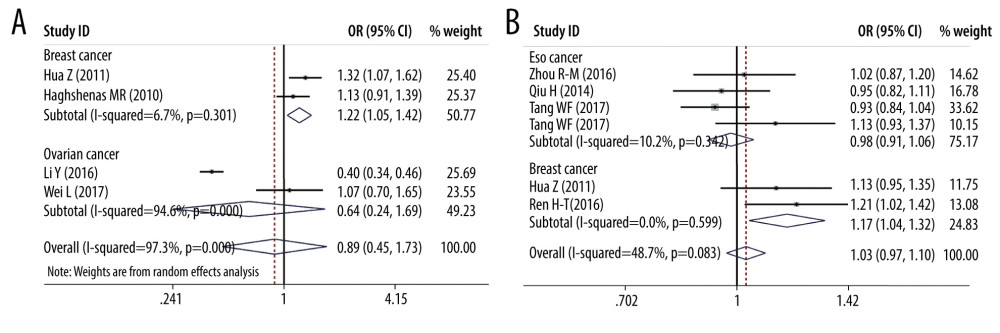 Figure 5. Forest plots of Subgroup analysis. (A) breast cancer (PD-1 rs2227981 in allele model); (B) breast cancer (PD-1 rs2227982 in allele model).
Figure 5. Forest plots of Subgroup analysis. (A) breast cancer (PD-1 rs2227981 in allele model); (B) breast cancer (PD-1 rs2227982 in allele model). 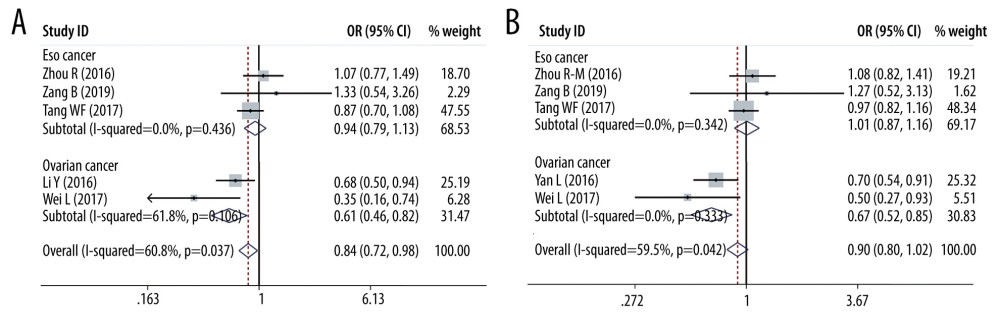 Figure 6. Forest plots of Subgroup analysis. (A) ovarian cancer (PD-1 rs36084323 in homozygote model); (B) ovarian cancer (PD-1 rs36084323 in homozygote model).
Figure 6. Forest plots of Subgroup analysis. (A) ovarian cancer (PD-1 rs36084323 in homozygote model); (B) ovarian cancer (PD-1 rs36084323 in homozygote model). 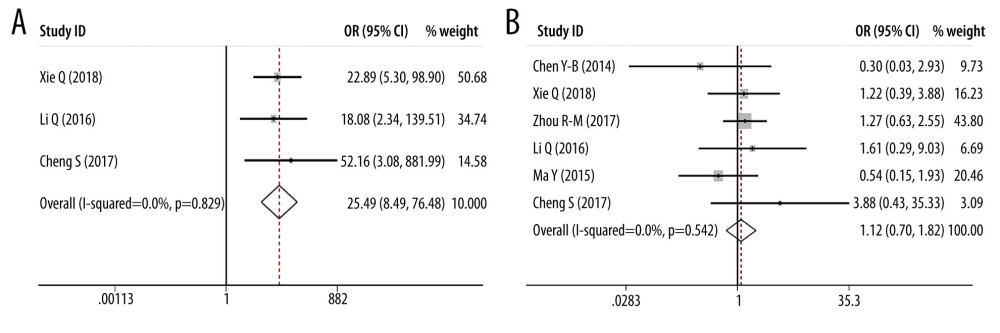 Figure 7. Forest plots of meta-analysis. (A) PD-L1 rs17718883 in homozygote model; (B) PD-L1 rs2890658 in homozygote model.
Figure 7. Forest plots of meta-analysis. (A) PD-L1 rs17718883 in homozygote model; (B) PD-L1 rs2890658 in homozygote model. 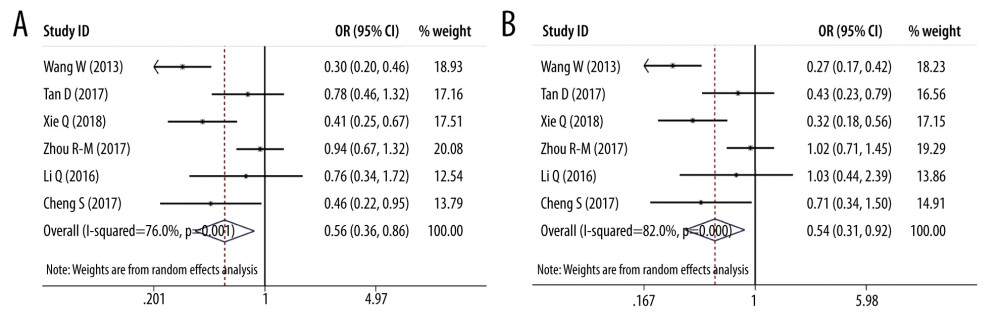 Figure 8. Forest plots of meta-analysis. (A) PD-L1 rs4143815 in heterozygote model; (B) PD-L1 rs4143815 in homozygote model.
Figure 8. Forest plots of meta-analysis. (A) PD-L1 rs4143815 in heterozygote model; (B) PD-L1 rs4143815 in homozygote model. 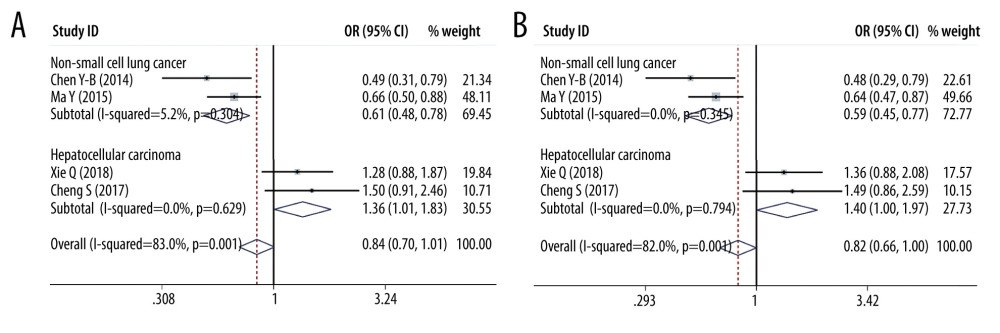 Figure 9. Forest plots of Subgroup analysis. (A) hepatocellular cancer (PD-L1 rs2890658 in allele model); (B) hepatocellular cancer (PD-L1 rs2890658 in recessive model).
Figure 9. Forest plots of Subgroup analysis. (A) hepatocellular cancer (PD-L1 rs2890658 in allele model); (B) hepatocellular cancer (PD-L1 rs2890658 in recessive model). 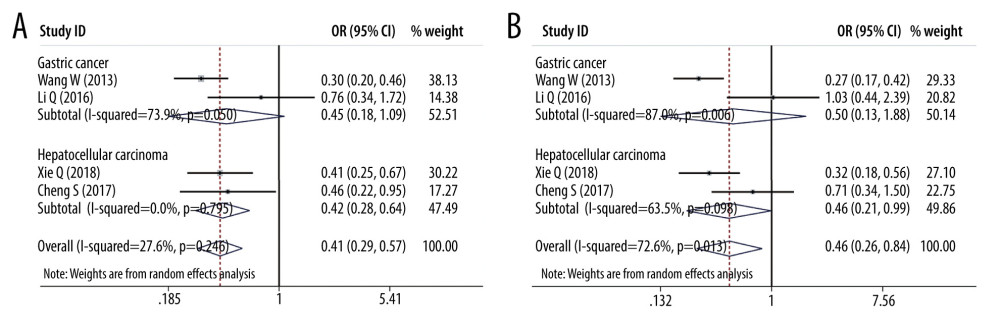 Figure 10. Forest plots of Subgroup analysis. (A) PD-L1 rs4143815 in heterozygote model (B) PD-L1 rs4143815 in homozygote model.
Figure 10. Forest plots of Subgroup analysis. (A) PD-L1 rs4143815 in heterozygote model (B) PD-L1 rs4143815 in homozygote model. 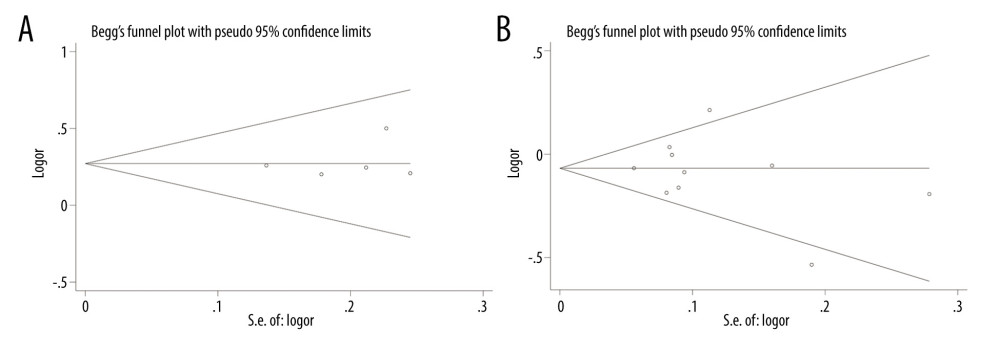 Figure 11. Publication bias. (A) Begg’s funnel plot for PD-1 rs11568821 in allele model; (B) Begg’s funnel plot for PD-1 rs36094323 in allele model.
Figure 11. Publication bias. (A) Begg’s funnel plot for PD-1 rs11568821 in allele model; (B) Begg’s funnel plot for PD-1 rs36094323 in allele model. 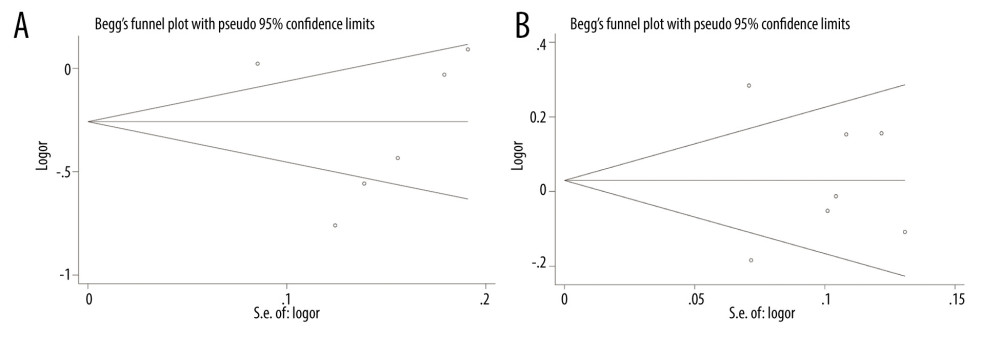 Figure 12. Publication bias. (A) Begg’s funnel plot for PD-L1 rs4143815; (B) Begg’s funnel plot for PD-1 rs7421861 in allele model.
Figure 12. Publication bias. (A) Begg’s funnel plot for PD-L1 rs4143815; (B) Begg’s funnel plot for PD-1 rs7421861 in allele model. Tables
Table 1. Characteristics of the selected articles.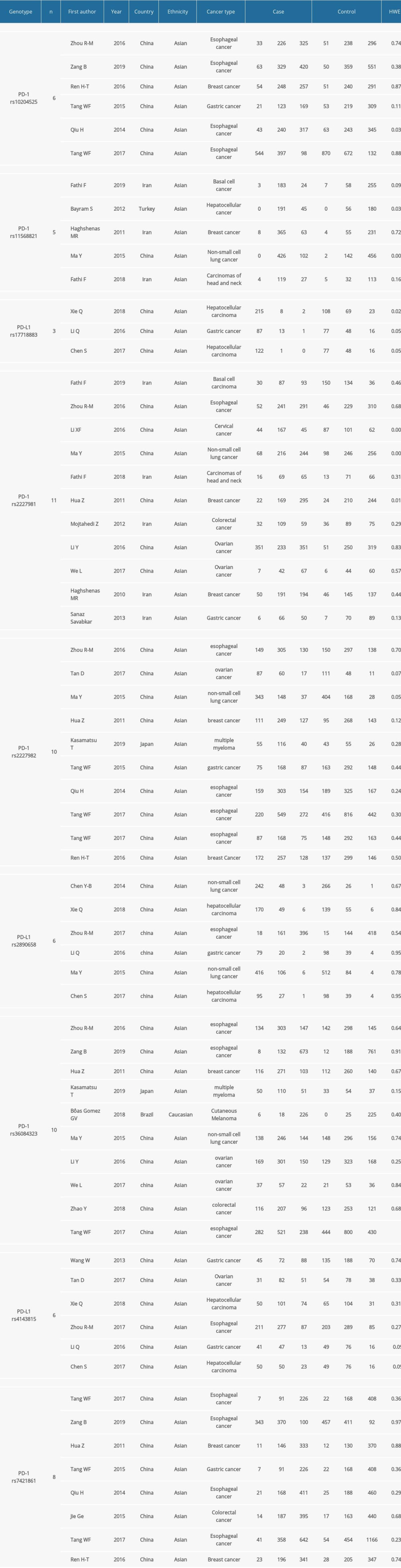 Table 2. Quality assessment based on the Newcastle-Ottawa Scale of trials included in this meta-analysis.
Table 2. Quality assessment based on the Newcastle-Ottawa Scale of trials included in this meta-analysis.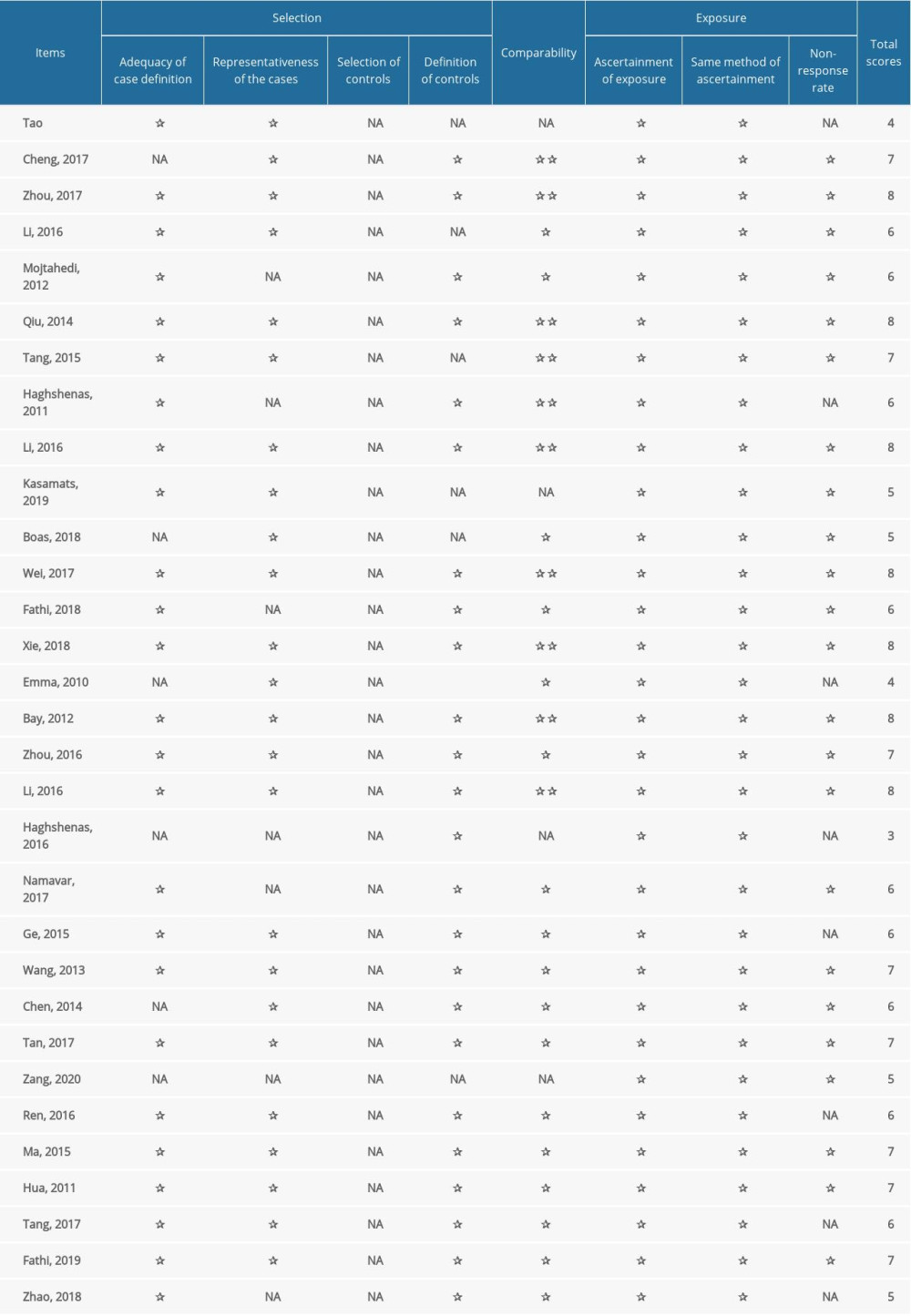 Table 3. Meta-analyses on PD-1 and PD-L1 variants and cancer susceptibility.
Table 3. Meta-analyses on PD-1 and PD-L1 variants and cancer susceptibility.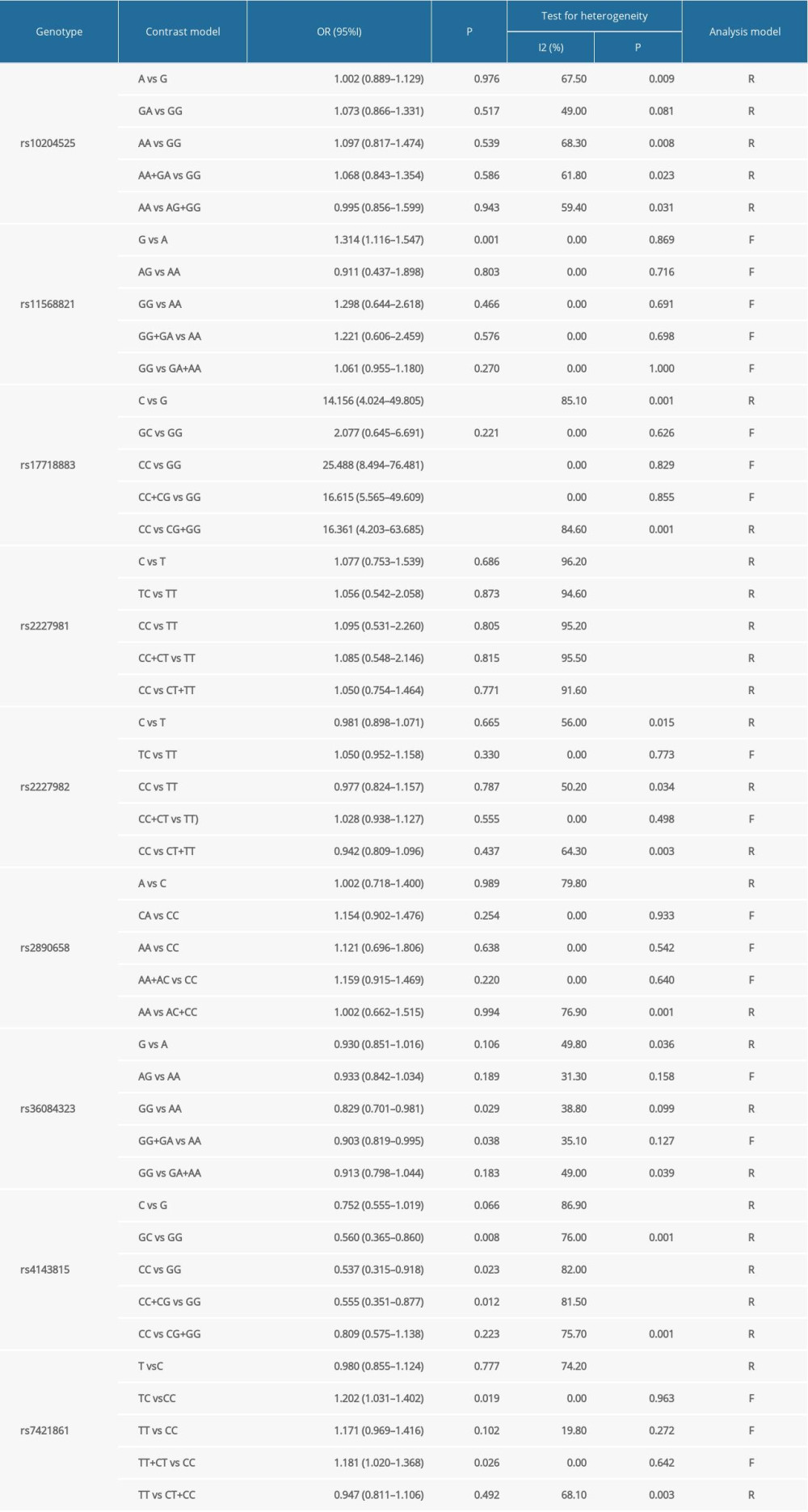 Table 4. Subgroup analyses based on cancer type.
Table 4. Subgroup analyses based on cancer type.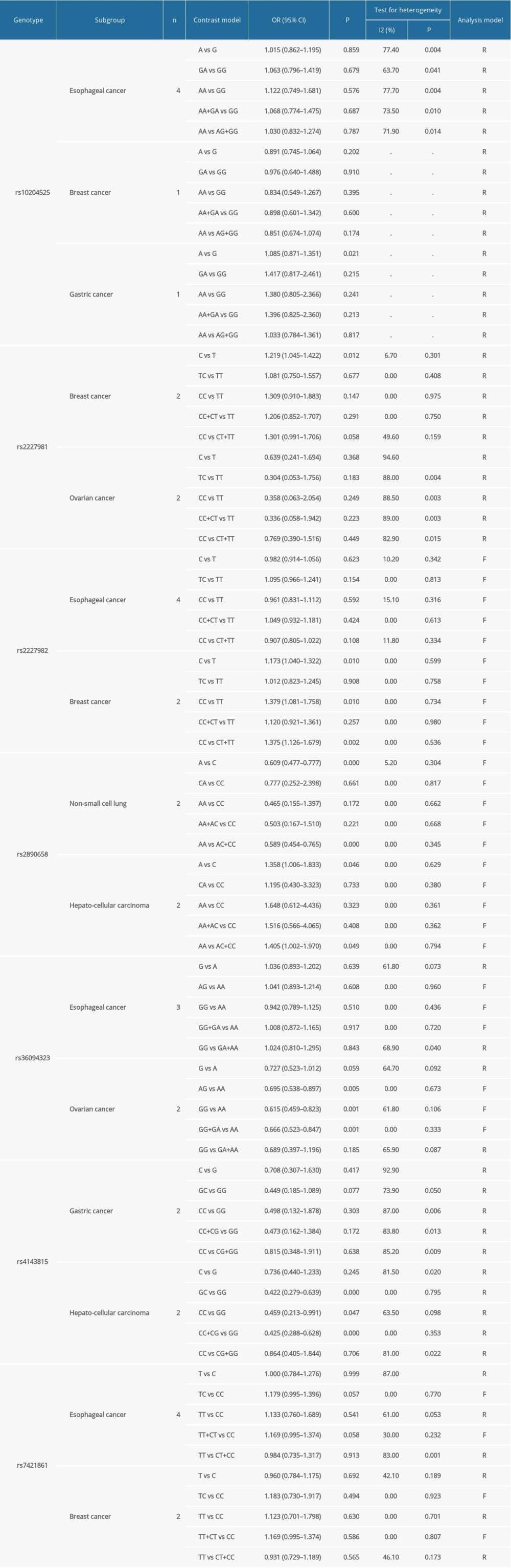 Table 5. Publication bias consequences.
Table 5. Publication bias consequences.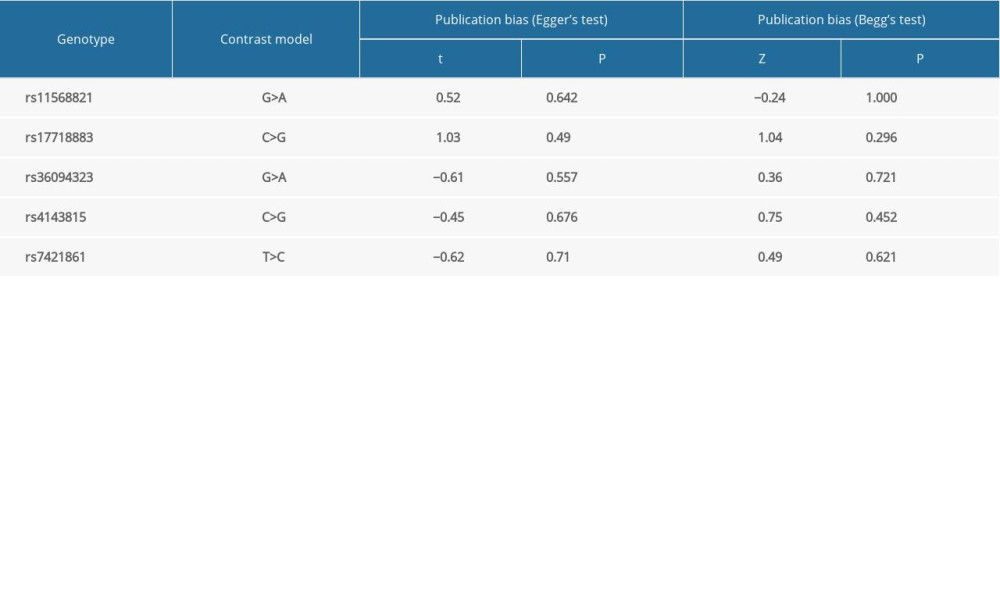
References
1. Siegel RL, Miller KD, Jemal A, Cancer statistics, 2020: Cancer J Clin, 2020; 70; 7-30
2. Yang Y, Cancer immunotherapy: Harnessing the immune system to battle cancer: J Clin Invest, 2015; 125; 3335-37
3. Salmaninejad A, Valilou SF, Shabgah AG, PD-1/PD-L1 pathway: Basic biology and role in cancer immunotherapy: J Cell Physiol, 2019; 234; 16824-37
4. Han Y, Liu D, Li L, PD-1/PD-L1 pathway: current researches in cancer: Am J Cancer Res, 2020; 10; 727-42
5. Gong J, Chehrazi-Raffle A, Reddi S, Salgia R, Development of PD-1 and PD-L1 inhibitors as a form of cancer immunotherapy: A comprehensive review of registration trials and future considerations: J Immunother Cancer, 2018; 6; 8
6. Gou Q, Dong C, Xu H, PD-L1 degradation pathway and immunotherapy for cancer: Cell Death Dis, 2020; 11; 955
7. Cha J, Chan L, Li C, Mechanisms controlling PD-L1 expression in cancer: Mol Cell, 2019; 76; 359-70
8. Guan J, Lim KS, Mekhail T, Chang C, Programmed death ligand-1 (PD-L1) expression in the programmed death receptor-1 (PD-1)/PD-L1 blockade: A key player against various cancers: Arch Pathol Lab Med, 2017; 141; 851-61
9. Ren H, Li Y, Wang X, PD-1 rs2227982 polymorphism is associated with the decreased risk of breast cancer in Northwest Chinese women: A hospital-based observational study: Medicine, 2016; 95; e3760
10. Da L, Zhang Y, Zhang C, The PD-1 rs36084323 A>G polymorphism decrease cancer risk in Asian: A meta-analysis: Pathol Res Pract, 2018; 214; 1758-64
11. Zang B, Chen C, Zhao J, PD-1 gene rs10204525 and rs7421861 polymorphisms are associated with increased risk and clinical features of esophageal cancer in a Chinese Han population: Aging, 2020; 12; 3771-90
12. Dong W, Gong M, Shi Z, Programmed cell death-1 polymorphisms decrease the cancer risk: A meta-analysis involving twelve case-control studies: PLoS One, 2016; 11; e0152448
13. Tan D, Sheng L, Yi Q, Correlation of PD-1/PD-L1 polymorphisms and expressions with clinicopathologic features and prognosis of ovarian cancer: Cancer Biomark, 2018; 21; 287-97
14. Zou J, Wu D, Li T, Association of PD-L1 gene rs4143815 C>G polymorphism and human cancer susceptibility: A systematic review and meta-analysis: Pathol Res Pract, 2019; 215; 229-34
15. Zhang J, Zhao T, Xu C, The association between polymorphisms in the PDCD1 gene and the risk of cancer: A PRISMA-compliant meta-analysis: Medicine, 2016; 95; e4423
16. Xie Q, Chen Z, Xia L, Correlations of PD-L1 gene polymorphisms with susceptibility and prognosis in hepatocellular carcinoma in a Chinese Han population: GENE, 2018; 674; 188-94
17. Zhao X, Peng Y, Li X, The association of PD-L1 gene polymorphisms with non-small-cell lung cancer susceptibility and clinical outcomes in a Chinese population: Int J Clin Exp Pathol, 2020; 13; 2130-36
18. Zhao Y, Gang M, Pang HAssociation of gene polymorphisms with colorectal cancer in chinese han population: Chinese Journal of Medical Genetics, 2018; 35; 219-23 [in Chinese]
19. Cheng SPD-L1 single nucleotide polymorphisms and serum detection in primary liver cancer and its clinical significance: Third Military Medical University, 2017 [in Chinese]
20. Mojtahedi Z, Mohmedi M, Rahimifar S, Programmed death-1 gene polymorphism (PD-1.5 C/T) is associated with colon cancer: Gene, 2012; 508; 229-32
21. Qiu H, Zheng L, Tang W, Programmed death-1 (PD-1) polymorphisms in Chinese patients with esophageal cancer: Clin Biochem, 2014; 47; 612-17
22. Tang W, Chen Y, Chen S, Programmed death-1 (PD-1) polymorphism is associated with gastric cardia adenocarcinoma: Int J Clin Exp Med, 2015; 8; 8086-93
23. Haghshenas MR, Naeimi S, Talei A, Program death 1 (PD1) haplotyping in patients with breast carcinoma: Mol Biol Rep, 2011; 38; 4205-10
24. Qing LiDiagnosis and prognostic value of PD-L1 expression and gene polymorphisms in gastric cancer: Third Military Medical University, 2016 [in Chinese]
25. Li Y, Zhang H, Kang S, The effect of polymorphisms in PD-1 gene on the risk of epithelial ovarian cancer and patients’ outcomes: Gynecol Oncol, 2017; 144; 140-45
26. Zhou R, Li Y, Liu J, Programmed death-1 ligand-1 gene rs2890658 polymorphism associated with the risk of esophageal squamous cell carcinoma in smokers: Cancer Biomark, 2017; 21; 65-71
27. Gomez GVB, Rinck-Junior JA, Oliveira C, PDCD1 gene polymorphisms as regulators of T-lymphocyte activity in cutaneous melanoma risk and prognosis: Pigment Cell Melanoma Res, 2018; 31; 308-17
28. Fathi F, Faghih Z, Khademi B, PD-1 haplotype combinations and susceptibility of patients to squamous cell carcinomas of head and neck: Immunol Invest, 2019; 48; 1-10
29. Bayram S, Akkız H, Ülger Y, Lack of an association of programmed cell death-1 PD1.3 polymorphism with risk of hepatocellular carcinoma susceptibility in Turkish population: A case-control study: Gene, 2012; 511; 308-13
30. Zhou R, Li Y, Wang N, Association of programmed death-1 polymorphisms with the risk and prognosis of esophageal squamous cell carcinoma: Cancer Genet, 2016; 209; 365-75
31. Li XF, Jiang XQ, Zhang JW, Jia YJ, Association of the programmed cell death-1 PD1.5 C>T polymorphism with cervical cancer risk in a Chinese population: Genet Mol Res, 2016; 15; 15016357
32. Wei L, Wang J, Men Y, Liao XThe relationship between PD-1 gene polymorphism and EOC susceptibility and prognosis: Study on maternal and child health in china, 2017; 28; 1608-12 [in Chinese]
33. Namavar JF, Samadi M, Mojtahedi Z, Association of PD-1.5 C/T, but not PD-1.3 G/A, with malignant and benign brain tumors in Iranian patients: Immunol Invest, 2017; 46; 469-80
34. Ge J, Zhu L, Zhou J, Association between co-inhibitory molecule gene tagging single nucleotide polymorphisms and the risk of colorectal cancer in Chinese: J Cancer Res Clin Oncol, 2015; 141; 1533-44
35. Chen YB, Mu CY, Chen C, Huang JA, Association between single nucleotide polymorphism of PD-L1 gene and non-small cell lung cancer susceptibility in a Chinese population: Asia Pac J Clin Oncol, 2014; 10; e1-6
36. Ma Y, Liu X, Zhu J, Polymorphisms of co-inhibitory molecules (CTLA-4/PD-1/PD-L1) and the risk of non-small cell lung cancer in a Chinese population: Int J Clin Exp Med, 2015; 8; 16585-91
37. Hua Z, Li D, Xiang G, PD-1 polymorphisms are associated with sporadic breast cancer in Chinese Han population of Northeast China: Breast Cancer Res Treat, 2011; 129; 195-201
38. Wang W, Li F, Mao Y, A miR-570 binding site polymorphism in the B7-H1 gene is associated with the risk of gastric adenocarcinoma: Hum Genet, 2013; 132; 641-48
39. Fathi F, Ebrahimi M, Eslami A, Association of programmed death-1 gene polymorphisms with the risk of basal cell carcinoma: Int J Immunogenet, 2019; 46; 444-50
40. Tang W, Chen S, Chen Y, Programmed death-1 polymorphisms is associated with risk of esophagogastric junction adenocarcinoma in the Chinese Han population: A case-control study involving 2,740 subjects: Oncotarget, 2017; 8; 39198-208
41. Kasamatsu T, Awata M, Ishihara R, PDCD1 and PDCD1LG1 polymorphisms affect the susceptibility to multiple myeloma: Clin Exp Med, 2020; 20; 51-62
42. Heinrichs S, Look AT, Identification of structural aberrations in cancer by SNP array analysis: Genome Biol, 2007; 8; 219
43. Qin W, Hu L, Zhang X, The diverse function of PD-1/PD-L pathway beyond cancer: Front Immunol, 2019; 10; 2298
44. Hashemi M, Karami S, Sarabandi S, Association between PD-1 and PD-L1 polymorphisms and the risk of cancer: A meta-analysis of case-control studies: Cancers, 2019; 11; 11081150
45. Topalian SL, Taube JM, Pardoll DM, Neoadjuvant checkpoint blockade for cancer immunotherapy: Science, 2020; 367(6477); aax0182
46. Akinleye A, Rasool Z, Immune checkpoint inhibitors of PD-L1 as cancer therapeutics: J Hematol Oncol, 2019; 12; 92
47. Goodman A, Patel SP, Kurzrock R, PD-1-PD-L1 immune-checkpoint blockade in B-cell lymphomas: Nat Rev Clin Oncol, 2017; 14; 203-20
48. Gomez S, Tabernacki T, Kobyra J, Combining epigenetic and immune therapy to overcome cancer resistance: Semin Cancer Biol, 2020; 65; 99-113
Figures
 Figure 1. Flowchart illustrating the search strategy for PD-1 and PD-L1 variants and cancer.
Figure 1. Flowchart illustrating the search strategy for PD-1 and PD-L1 variants and cancer. Figure 2. Forest plots of meta-analysis. (A) PD-1 rs11568821 in allele model (B) PD-1 rs10204525 in allele model.
Figure 2. Forest plots of meta-analysis. (A) PD-1 rs11568821 in allele model (B) PD-1 rs10204525 in allele model. Figure 3. Forest plots of meta-analysis. (A) PD-1 rs36084323 in dominant model (B) PD-1 rs2227981 in dominant model.
Figure 3. Forest plots of meta-analysis. (A) PD-1 rs36084323 in dominant model (B) PD-1 rs2227981 in dominant model. Figure 4. Forest plots of meta-analysis. (A) PD-1 rs7421861 in heterozygote model (B) PD-1 rs7421861 in dominant model.
Figure 4. Forest plots of meta-analysis. (A) PD-1 rs7421861 in heterozygote model (B) PD-1 rs7421861 in dominant model. Figure 5. Forest plots of Subgroup analysis. (A) breast cancer (PD-1 rs2227981 in allele model); (B) breast cancer (PD-1 rs2227982 in allele model).
Figure 5. Forest plots of Subgroup analysis. (A) breast cancer (PD-1 rs2227981 in allele model); (B) breast cancer (PD-1 rs2227982 in allele model). Figure 6. Forest plots of Subgroup analysis. (A) ovarian cancer (PD-1 rs36084323 in homozygote model); (B) ovarian cancer (PD-1 rs36084323 in homozygote model).
Figure 6. Forest plots of Subgroup analysis. (A) ovarian cancer (PD-1 rs36084323 in homozygote model); (B) ovarian cancer (PD-1 rs36084323 in homozygote model). Figure 7. Forest plots of meta-analysis. (A) PD-L1 rs17718883 in homozygote model; (B) PD-L1 rs2890658 in homozygote model.
Figure 7. Forest plots of meta-analysis. (A) PD-L1 rs17718883 in homozygote model; (B) PD-L1 rs2890658 in homozygote model. Figure 8. Forest plots of meta-analysis. (A) PD-L1 rs4143815 in heterozygote model; (B) PD-L1 rs4143815 in homozygote model.
Figure 8. Forest plots of meta-analysis. (A) PD-L1 rs4143815 in heterozygote model; (B) PD-L1 rs4143815 in homozygote model. Figure 9. Forest plots of Subgroup analysis. (A) hepatocellular cancer (PD-L1 rs2890658 in allele model); (B) hepatocellular cancer (PD-L1 rs2890658 in recessive model).
Figure 9. Forest plots of Subgroup analysis. (A) hepatocellular cancer (PD-L1 rs2890658 in allele model); (B) hepatocellular cancer (PD-L1 rs2890658 in recessive model). Figure 10. Forest plots of Subgroup analysis. (A) PD-L1 rs4143815 in heterozygote model (B) PD-L1 rs4143815 in homozygote model.
Figure 10. Forest plots of Subgroup analysis. (A) PD-L1 rs4143815 in heterozygote model (B) PD-L1 rs4143815 in homozygote model. Figure 11. Publication bias. (A) Begg’s funnel plot for PD-1 rs11568821 in allele model; (B) Begg’s funnel plot for PD-1 rs36094323 in allele model.
Figure 11. Publication bias. (A) Begg’s funnel plot for PD-1 rs11568821 in allele model; (B) Begg’s funnel plot for PD-1 rs36094323 in allele model. Figure 12. Publication bias. (A) Begg’s funnel plot for PD-L1 rs4143815; (B) Begg’s funnel plot for PD-1 rs7421861 in allele model.
Figure 12. Publication bias. (A) Begg’s funnel plot for PD-L1 rs4143815; (B) Begg’s funnel plot for PD-1 rs7421861 in allele model. Tables
 Table 1. Characteristics of the selected articles.
Table 1. Characteristics of the selected articles. Table 2. Quality assessment based on the Newcastle-Ottawa Scale of trials included in this meta-analysis.
Table 2. Quality assessment based on the Newcastle-Ottawa Scale of trials included in this meta-analysis. Table 3. Meta-analyses on PD-1 and PD-L1 variants and cancer susceptibility.
Table 3. Meta-analyses on PD-1 and PD-L1 variants and cancer susceptibility. Table 4. Subgroup analyses based on cancer type.
Table 4. Subgroup analyses based on cancer type. Table 5. Publication bias consequences.
Table 5. Publication bias consequences. Table 1. Characteristics of the selected articles.
Table 1. Characteristics of the selected articles. Table 2. Quality assessment based on the Newcastle-Ottawa Scale of trials included in this meta-analysis.
Table 2. Quality assessment based on the Newcastle-Ottawa Scale of trials included in this meta-analysis. Table 3. Meta-analyses on PD-1 and PD-L1 variants and cancer susceptibility.
Table 3. Meta-analyses on PD-1 and PD-L1 variants and cancer susceptibility. Table 4. Subgroup analyses based on cancer type.
Table 4. Subgroup analyses based on cancer type. Table 5. Publication bias consequences.
Table 5. Publication bias consequences. In Press
15 Apr 2024 : Laboratory Research
The Role of Copper-Induced M2 Macrophage Polarization in Protecting Cartilage Matrix in OsteoarthritisMed Sci Monit In Press; DOI: 10.12659/MSM.943738
07 Mar 2024 : Clinical Research
Knowledge of and Attitudes Toward Clinical Trials: A Questionnaire-Based Study of 179 Male Third- and Fourt...Med Sci Monit In Press; DOI: 10.12659/MSM.943468
08 Mar 2024 : Animal Research
Modification of Experimental Model of Necrotizing Enterocolitis (NEC) in Rat Pups by Single Exposure to Hyp...Med Sci Monit In Press; DOI: 10.12659/MSM.943443
18 Apr 2024 : Clinical Research
Comparative Analysis of Open and Closed Sphincterotomy for the Treatment of Chronic Anal Fissure: Safety an...Med Sci Monit In Press; DOI: 10.12659/MSM.944127
Most Viewed Current Articles
17 Jan 2024 : Review article
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
14 Dec 2022 : Clinical Research
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387
01 Jan 2022 : Editorial
Editorial: Current Status of Oral Antiviral Drug Treatments for SARS-CoV-2 Infection in Non-Hospitalized Pa...DOI :10.12659/MSM.935952
Med Sci Monit 2022; 28:e935952








