11 July 2021: Database Analysis
Identification of as an Immune-Related Biomarker Associated with Tumorigenesis and Prognosis in Cutaneous Melanoma Patients
Zebing Si1DEF*, Honghe Hu1AGDOI: 10.12659/MSM.932052
Med Sci Monit 2021; 27:e932052
Abstract
BACKGROUND: Melanoma is one of the most lethal tumors and its treatment is still challenging. It is urgent to detect novel therapy targets in melanoma.
MATERIAL AND METHODS: The GEO dataset was used to obtain a list of DEGS (differentially-expressed genes). Integrative bioinformatics analyses, including HPRD database, TCGA data, and TIMER, were performed to determine the role of CXCL13 in SKCM (skin cutaneous melanoma) progression and the immune environment. Furthermore, Pearson correlation coefficient analysis was used to measure correlations between CXCL13 and its co-expressed genes. Survival analysis, GO, and KEGG enrichment analysis were performed to investigate the role of CXCL13 in SKCM.
RESULTS: A total of 41 DEGs were identified in 3 GEO datasets, and 4 out of 41 DEGs are hub genes. Among the 4 hub genes, CXCL13 is involved in the most KEGG terms. CXCL13 is co-expressed with well-known immune checkpoint blockade targets, and it was associated with better overall survival. In addition, CXCL13 levels in infiltrating immune cells (neutrophil and myeloid dendritic cells) affect prognosis and survival in SKCM. Functional enrichment analysis clarified that CXCL13-co-expressed top 30 genes were associated with immune signaling pathways. Network analysis identified CXCL13 as a hub gene that interacts with CXCR5 to participate in immune-related biological process.
CONCLUSIONS: This study found that CXCL13 is associated with SKCM tumorigenesis and prognosis and immune infiltrations. Our result suggests that CXCL13 has great potential in development of novel immunotherapy targets in melanoma.
Keywords: CXCL13 Protein, Human, Melanoma, Biomarkers, Tumor, carcinogenesis, Chemokine CXCL13, Computational Biology, Databases, Genetic, Skin Neoplasms, tumor microenvironment
Background
Melanoma is a malignant tumor that causes approximately three-quarters of skin cancer-related deaths [1]. Melanoma treatment has been revolutionized with the approval of BRAFV600E inhibitors, which can significantly improve response and overall survival for patients with the BRAFV600E mutation [2]. However, patients develop acquired resistance to BRAFV600E inhibitors treatment [3]. Subsequent development of targeted immune checkpoints therapy has prolonged the survival of patients, such as with targeted CTLA-4 (ipilimumab, approved in 2011), and PD-1 (nivolumab, pembrolizumab, approved in 2014) has extended recurrence-free survival in high-risk resected melanoma patients [4,5]. However, melanoma recurrence was approximately 25–30% within 1 year in patients who were targeted by PD1, and PD1 monotherapy was not effective in those patients [5]. To overcome the drug resistance, it is urgent to explore novel therapeutic targets and treatment strategies for melanoma.
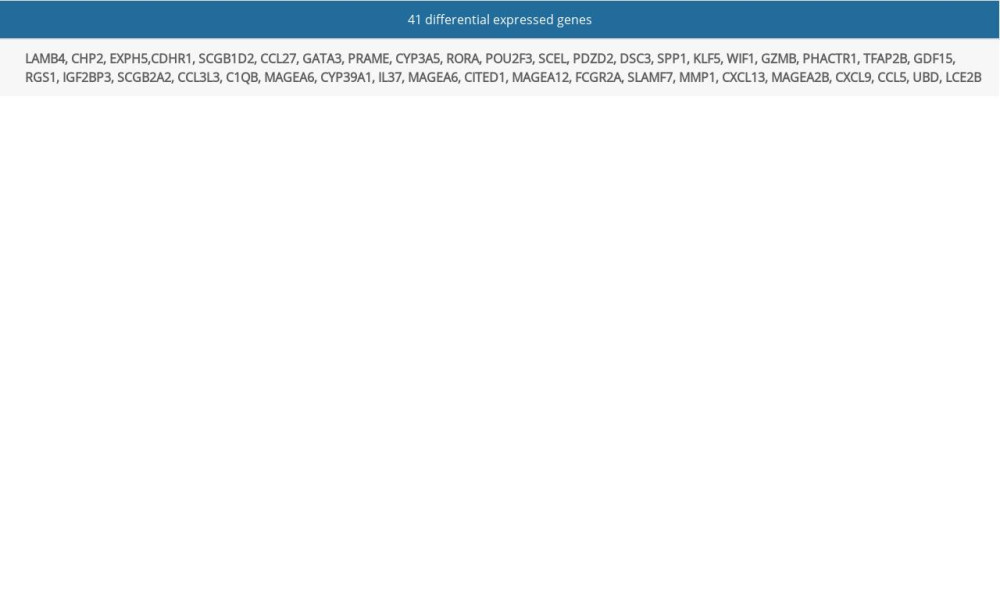
In our study, based on 3 datasets from the GEO database, we found that 41 differentially-expressed genes (DEGs) were overexpressed in melanoma compared to normal samples. Through network analysis, 4 (CXCL13, MMP1, SPP1, GZMB) out of the 41 DEGs had a high degree. KEGG enrichment analysis for the 41 DEGs indicated that CXCL1 is involved in 3 pathways. Moreover, Kaplan-Meier survival analysis showed that high
Material and Methods
IDENTIFICATION OF DEGS:
We accessed GEO database to obtain the expression data. GSE15605 (58 melanoma samples and 16 normal samples), GSE46517 (104 melanoma samples and 12 normal samples), and GSE114445 (16 melanoma samples and 18 normal samples) were adopted to perform the differentially expression. The DEGs between melanoma and normal samples were collected using GEO2R (
NETWORK CONSTRUCTION AND FUNCTIONAL GENE SETS:
The network is an undirected graph G (V, E) where V stands for the genes and edges (i, j) ∈ E are weighted by PCC. Protein interaction networks were sourced from HPRD (http://www.hprd.org) database [7], the version is release 9. Protein self-interactions were removed, resulting in 39240 interactions among 9616 proteins. Cytoscape can create network interaction maps based on input genes and gene interaction [8]. The HPRD was used to extract the protein interaction of DEGs. 41DEGs were mapped to the HPRD, and the interaction was plotted by Cytoscape software (version 3.8.2, https://cytoscape.org/). The degree of each DEG was defined as the number of each DEG connected directly to its neighbor in HPRD. Only the nodes with degree greater than 5 were regarded as high-degree nodes.
SURVIVAL ANALYSIS:
The gene expression RNAseq and the survival time of TCGA-SKCM were downloaded from UCSC Xena (
Survival curves were estimated using the Kaplan-Meier method and compared across groups with the log-rank test. The
CO-EXPRESSION ANALYSIS:
Gene and gene co-expression were analyzed statistically using Pearson’s correlation coefficient (PCC). The sample size of the gene expression was 473 melanoma samples. The scatter plots of 3 known immune genes (PD1, PDL1 and CTLA4) correlated with CXCL13 were plotted using ggplot2, ggpubr, and ggpmisc packages in R (version 3.6). The heat map of the top 30 highly co-expressed genes with CXCL13 was plotted using the pheatmap package in R (version 3.6,
ENRICHMENT ANALYSIS:
We then performed GO and KEGG enrichment analyses of the top 30 high-correction-associated genes using the “clusterProfiler” package in R (version 3.6,
TIMER:
TIMER (http://timer.cistrome.org) is a comprehensive resource for systematic analysis (immune association, cancer exploration, and immune estimation) of immune infiltrates across diverse cancer types [9]. The association between immune infiltrates (CD4 T cells, neutrophils, macrophages, and myeloid dendritic cells) and clinical outcome for CXCL13 was explored using the TIMER online service.
Results
IDENTIFICATION OF DIFFERENTIAL GENES:
To find the potential and reliable target genes, we first extracted genes which were differentially expressed in melanoma and normal samples. Through differential expression analysis, each of these datasets (GSE15065, GSE46517, and GSE114445) produced the corresponding list of DEGs. The 41 overlapped genes (Supplementary Table 1), as shown in the Venn diagram, are common DEGs in the 3 datasets (Figure 1A)
KEGG ENRICHMENT ANALYSIS OF THE DEGS:
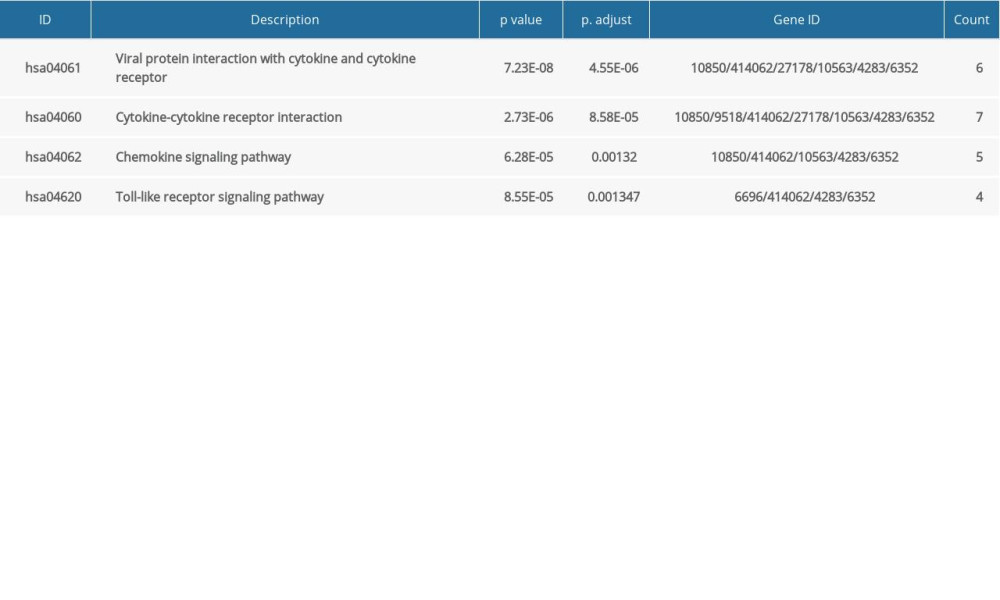
KEGG enrichment of 41 DEGs was performed using the clusterProfiler package in R software, then displayed in bubble charts (P<0.05, Figure 1B). The size of the bubble represents the number of genes in the KEGG terms, and the color of bubbles represents the adjusted P value in the KEGG terms. KEGG enrichment analysis indicated that changes in biologic pathways were significantly enriched in cytokine-cytokine receptor interaction, chemokine signaling pathway, and Toll-like receptor signaling pathway. These results revealed that immune response and inflammatory response play important roles in melanoma tumorigenesis.
PPI NETWORK CONSTRUCTION AND HUB GENES IDENTIFICATION:
We mapped 41 DEGs to the HPRD database and extracted their directly connected proteins, including 113 nodes (gene) and 92 edges (gene and gene interaction). Then, we input the information of 92 edges into Cytoscape. In Figure 2A, we display the high-degree genes-associated network. Figure 2A shows 4 (MMP1, SPP1, CXCL13, and GZMB) hub genes (the node with yellow color) with a degree greater than 5. Combined with the KEGG enrichment information of 41 DEGs, and among the 4 hub genes, CXC13 was found to be involved in 3 KEGG pathway terms (Supplementary Table 2).
Figure 2A shows that CXCL13 is connected with CXCR5. To investigate whether CXCL13 is co-expressed with CXCR5, we calculated the Pearson correlation coefficient. There was a positive correlation between CXCL13 expression and CXCR5 expression (PCC=0.74). High CXCL13 and CXCR5 expression were significantly related to longer survival time in SKCM patients (P=0.001). CXCL13 can interact with its ligand CXCR5 and play vital roles in apoptosis, proliferation, and differentiation of immune cells. It has been revealed that CXCL13 makes pivotal contributions to the biological process of multiple cancers via CXCL13/CXCR5 signaling [10]. For SKCM, in a previous study, Huang et al found that CXCL13 acts as an immune-related biomarker [11], which is in agreement with our results. The prognostic value and biological function of CXCL13 in SKCM are unclear and need to be better characterized. Here, we screened out available datasets associated with SKCM from public databases to systematically investigate the mechanism of CXCL13 on the progression, prognosis, and microenvironment of SKCM.
CXCL13 LEVEL IN PATIENTS WITH SKCM:
Early detection and treatment of SKCM is still a challenge that perplexes clinicians. Therefore, new therapy targets are needed to improve treatment efficacy. We first detected and compared the mRNA level of
THE RELATIONSHIP BETWEEN CXCL13 EXPRESSION AND CLINICAL OUTCOMES OF SKCM PATIENTS:
We also evaluated the prognostic value of CXCL13 in SKCM from TCGA. Kaplan-Meier survival curves of OS were produced utilizing CXCL13 expression data and survival information of SKCM. The sample with CXCL13 expression greater than the 50th percentile of CXCL13 expression was treated as high CXCL13 expression, and otherwise was treated as low CXCL13 expression. The survival analysis showed that low expression of CXCL13 (P =0.00037) predicted poor OS (Figure 2B). In the above analysis, CXCL13 had lower expression in SKCM than in normal samples. This supports that CXCL13 low expression is related to poor OS. Therefore, it is conceivable that high CXCL13 expression is an independent risk factor and leads to a better prognosis in melanoma patients.
CXCL13 CO-EXPRESSION GENES ARE INVOLVED IN IMMUNE PATHWAYS:
We then focused on the function of the top 30 highly co-expressed genes with CXCL13 using GO enrichment analysis, showing that co-expression genes were correlated with CXCL13 in SKCM. PCC was utilized to analyze CXCL13 co-expressed genes from TCGA data. As shown in Figure 3A, the heat map displays the genes that showed significant positive correlations with CXCL13. The X axis represents SKCM samples, and the Y axis represents the rank of genes which are positive correlations with CXCL13. To explore the function of the top 30 highly co-expression genes, GO enrichment analyses were performed. LCK is a kinase that is among the top 30 highly co-expressed genes. Kinases play a crucial role in cancer formation and progress by modulating cancer cell migration, progression, and apoptosis [12]. The results of our study indicated that LCK is the vital regulator of CXCL13 in SKCM.
Figure 3B displays the top 10 most statistically significant GO terms. Biological processes (BP) were significantly enriched in T cell activation, lymphocyte differentiation, regulation of lymphocyte activation, regulation of T cell action, and regulation of cell-cell adhesion, which suggests CXCL13 has an effect on the transcriptome. Therefore, we found that the biological functions of these co-expressed genes are primarily related to immune-related pathways. Previous studies have demonstrated that the immune microenvironment plays key roles in immune evasion and metastasis of various cancers [13]. CXCL13 interacts with other genes to regulate the tumor immune environment, and CXCL13 co-expression genes make a crucial contribution to immune-related pathways. In summary, CXCL13 makes an important contribution to immune biological process.
CXCL13 IS CO-EXPRESSED WITH 3 WELL-KNOWN IMMUNE CHECKPOINT BLOCKADE TARGETS:
To assess the potential of
To explore the biological effects of CXCL13 in SKCM, we examined CXCL13 co-expression in SKCM. CXCL13 is co-expressed with CTLA4, PD-1 (PDCD1), and PD-L1 (CD274) in SKCM and the relationship is statically significant (Figure 4). Immune checkpoint therapies, including PD-1 or PD-L1, and CTLA4, have extended patient survival times in multiple cancer types. In Figure 4A, X represents the CXCL13 expression data across 473 SKCM samples and the Y represents the PDCD1 expression data across 473 SKCM samples. We found that CXCL13 is co-expressed with these immune checkpoint blockade genes. CXCL13 expression showed a strong positive association with expression of PDCD1 (PCC=0.87, P<0.01), PDL1 (PCC=0.66, P<0.01), and CTLA4 (PCC=0.57, P<0.01). This suggests CXCL13 interacts with immune checkpoint blockade genes and plays a vital in immune function. The positive correlation between the expression of these proteins and CXCL13 expression indicates that CXCL13 could be a predictor for active response to anti-PD-1/PD-L1 immunotherapy in SKCM since PD-L1 expression is a biomarker for predicting the anti-PD-1/PD-L1 immunotherapy response in cancer. The cooperative anticancer effects of anti-CD137 with anti-CTLA4 or/and -PD-1 antibodies in several cancer types exceed that of a single antibody, which shows the advantage of antibody combinations in the immune checkpoint therapies in cancer treatment. PD-L1 expression can act as a indictor for the anti-PD-1/PD-L1 immunotherapy response in cancer [14]. The positive correlation between the expression of PD-1/PD-L1/CTLA4 and CXCL13 expression indicates that CXCL13 upregulation could be a biomarker for the response to anti-PD-1/PD-L1/CTLA4 immunotherapy in SKCM.
To investigate whether the CXCL13 and the immune checkpoint gene have an effect on survival time, we performed survival analysis. We used the 50th quantile as a threshold to create high-expression and low-expression groups. Figure 5A shows the correlation between CXCL13 expression in SKCM expression data, CTLA4 expression in SKCM expression data, and survival time. Patients with high CXCL13 gene expression and high PD-1 gene expression had a longer survival time in SKCM (p=0.0011). In Figure 5B, patients with high CXCL13 gene expression and high PD-L1 gene expression had a longer survival time in SKCM (P<0.0001). In Figure 5C, patients with high CXCL13 gene expression and high CTLA4 gene expression had a longer survival time in SKCM (P<0.0001). This suggests that the efficacy of the combination of CXCL13 and PD-1/PD-L1/CTLA4 antibodies to treat SKCM. Thus, more investigations are urgently needed on the combination of CXCL13 and other antibodies to advance treatment effects and determine the underlying mechanisms.
MODULES EXPLORING THE ASSOCIATION BETWEEN IMMUNE INFILTRATES AND CXCL13 EXPRESSION OR CLINICAL UUTCOME IN TCGA:
CXCL13 mediates the migration and localization of immune cells. Increasing information shows that immune cell infiltration can accelerate tumor progression and recurrence and affect immunotherapy and clinical outcome. Correlation between CXCL13 in SKCM expression, abundance of immune infiltrates (neutrophil cells, macrophages, myeloid dendritic cells and CD4+ T cells), and survival time are shown in Figure 6. Patients with high CXCL13 gene expression and high neutrophil infiltrates had a longer survival time than patients with high gene expression and low neutrophil infiltrates (P<0.01). Patients with high CXCL13 gene expression and low macrophage infiltrates tended to have a longer survival time than those with high CXCL13 gene expression and high macrophage infiltrates (P<0.01). Patients with high CXCL13 gene expression and high myeloid dendritic cell infiltrates tended to a have longer survival times (P=0.003). Patients with low CXCL13 gene expression and low macrophage infiltrates tended to have a longer survival time than those with low CXCL13 gene expression and high macrophage infiltrates (P=0.02). CD4+ T cells can inhibit cancer growth via activating M1 macrophages. Patients with low CXCL13 gene expression and low T cell CD4+ infiltrates had a longer survival time than patients with low gene expression and high T cell CD4+ infiltrates (P=0.04). Thus, cumulative curve analysis showed that immune infiltrates were significantly associated with CXCL13 in SKCM, indicating that immune infiltrate cells significantly affect the prognosis. In this study, we found a significant correlation between the survival time and the expression of CXCL13 in the infiltration of the 4 immune cell types – neutrophils, CD4+ T cells, macrophages, and myeloid dendritic cells – indicating that CXCL13 expression in combination with immune cell status can predict prognosis. Further research is needed to explore the potential immune therapy using CXCL13.
Discussion
Melanoma is a malignant tumor. Although PD1 monotherapy and BRAF inhibitors were widely used in patients with melanoma, these therapies were not active in those patients with drug resistance. Therefore, it is urgent to detect more therapeutic targets and prognostic biomarkers.
In our study, 41 common DEGs were found in 3 datasets. KEGG enrichment analysis indicated that these DEGs play a crucial role in immune response and inflammatory response. After inputting 41 DEGs and their PPI into Cytoscape software, we obtained 4 (MMP1, SPP1, CXCL13, and GZMB) hub genes with a high degree. KEGG enrichment revealed that CXCL13 was enriched in the most KEGG pathway terms. Recent studies have reported that chemokine receptors expressed on melanoma and immune cells are closely associated with the prognosis and the efficacy of melanoma immunotherapy [15,16].
Our analysis suggests that
In addition, the expressions of CXCL13 were positively correlated with infiltration of 4 immune cell (neutrophils, CD4+ T cells, macrophages, and myeloid dendritic cells). Previous research reported that the degree of immune cell infiltration was related to favorable outcomes [20]. This is similar to our results that increasing levels of neutrophils and myeloid dendritic cells have a relationship with longer survival time in TCGA-SKCM patients.
Our results provide insights for future research on immunotherapy of SKCM.
It also could be highly useful in the identification of comorbidity patterns between diseases. Bioinformatics tools are novel alternatives for conventional approaches to get these data and integrate them in a biologically meaningful manner. Exploiting the bioinformatics tools, we have introduced
Conclusions
In summary, we systematically analyzed CXCL13 expression, prognosis, biological function, immune infiltration, and its relationship with 3 well-known immune checkpoint blockade targets. CXCL13 expression was closely related to survival rates of SKCM patients. CXCL13 can interact with other genes to regulate the immune pathways. It also has a correction with the expression of PD-1/PD-L1/CTLA4. In addition, CXCL13 is associated with infiltration of multiple immune cells. Research shows that CXCL13 may serve as potential therapeutic target in cutaneous melanoma, and it could be a biomarker for the response to anti-PD-1/PD-L1/CTLA4 immunotherapy in SKCM.
Figures
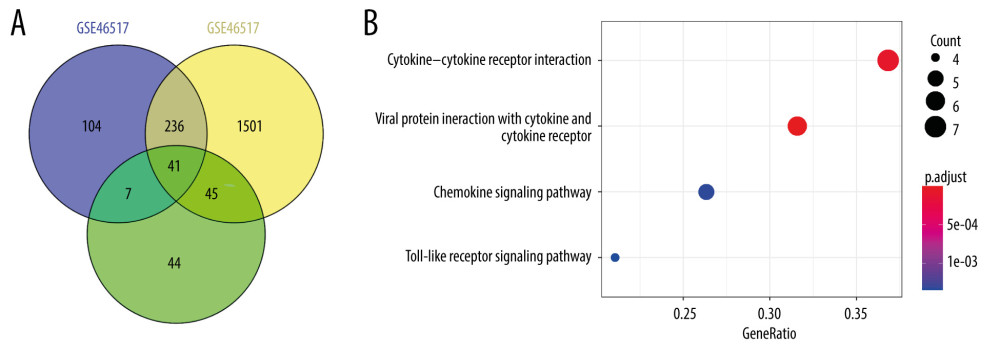 Figure 1. Venn diagram and the most significant KEGG pathways of 41DEGs. (A) The Venn diagram displays the number of DEGs in 3 datasets from the GEO database. (B) Four significant KEGG pathways of 41 DEGs. Adjusted p value <0.01. Figure 1A was produced using the Venny website (https://bioinfogp.cnb.csic.es/tools/venny/index.html, version 2.1). Figure 1B was produced using clusterProfiler packages in R (version 3.6, https://www.r-project.org/).
Figure 1. Venn diagram and the most significant KEGG pathways of 41DEGs. (A) The Venn diagram displays the number of DEGs in 3 datasets from the GEO database. (B) Four significant KEGG pathways of 41 DEGs. Adjusted p value <0.01. Figure 1A was produced using the Venny website (https://bioinfogp.cnb.csic.es/tools/venny/index.html, version 2.1). Figure 1B was produced using clusterProfiler packages in R (version 3.6, https://www.r-project.org/). 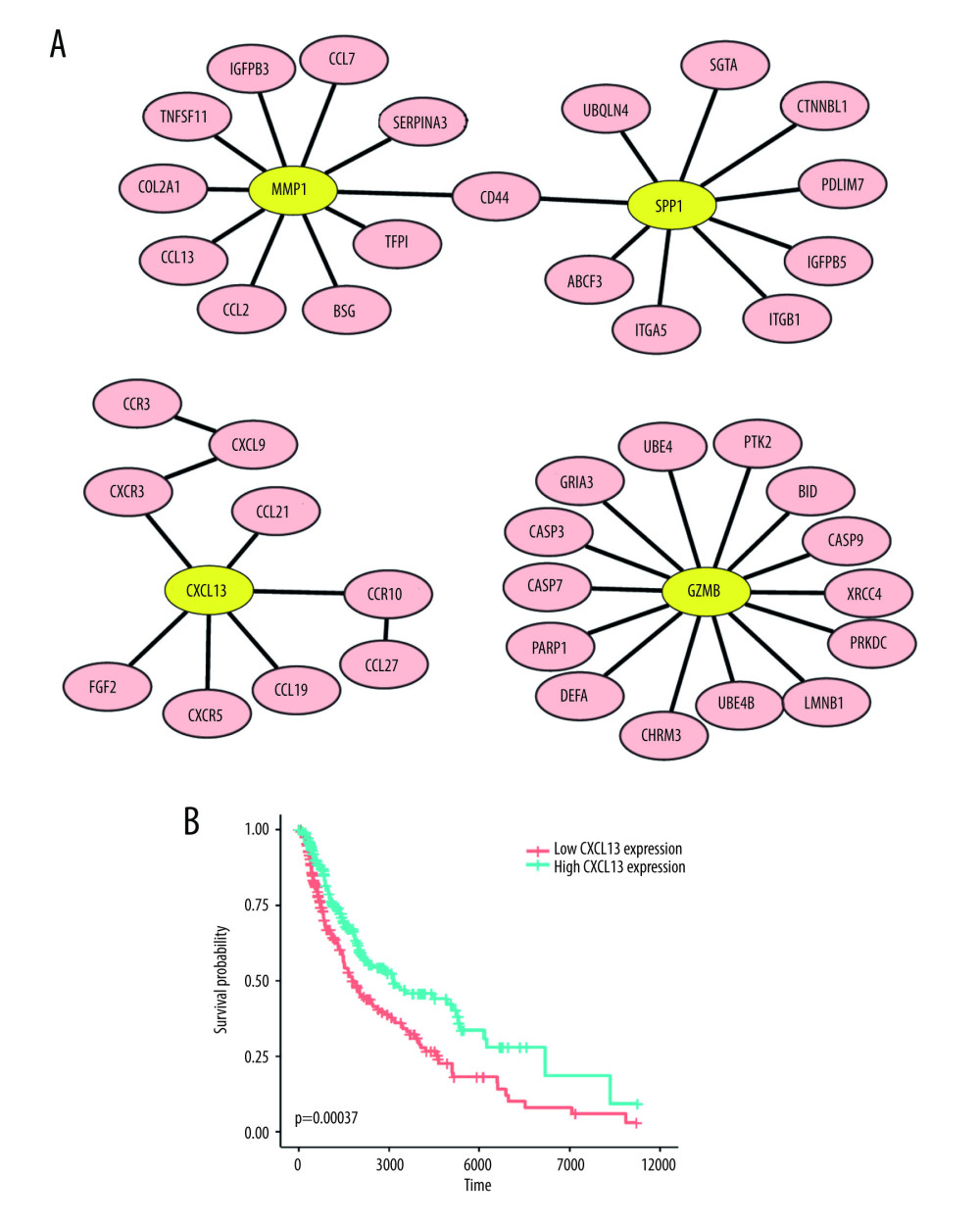 Figure 2. The network and survival plot. (A) The PPI (Protein-Protein Interaction) network of 4 high-degree DEGs (differentially-expressed genes). (B) Kaplan-Meier survival curves show that the expression of CXCL13 is consistently associated with better overall survival (OS) in SKCM. Figure 2A was produced using Cytoscape (version 3.8.2, https://cytoscape.org/). Figure 2B was produced using survival and survminer packages in R (version 3.6, https://www.r-project.org/).
Figure 2. The network and survival plot. (A) The PPI (Protein-Protein Interaction) network of 4 high-degree DEGs (differentially-expressed genes). (B) Kaplan-Meier survival curves show that the expression of CXCL13 is consistently associated with better overall survival (OS) in SKCM. Figure 2A was produced using Cytoscape (version 3.8.2, https://cytoscape.org/). Figure 2B was produced using survival and survminer packages in R (version 3.6, https://www.r-project.org/). 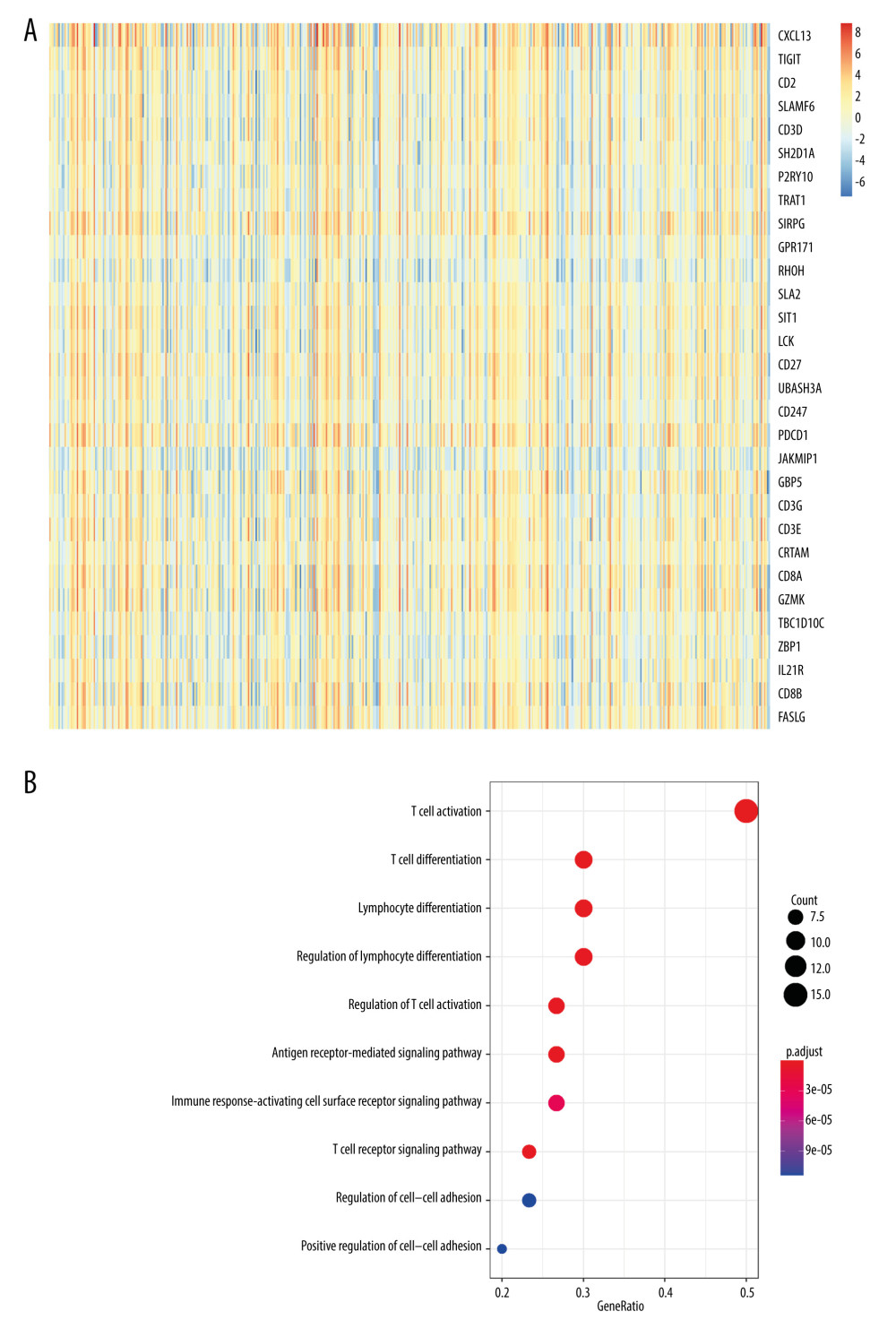 Figure 3. CXCL13 associated top 30 co-expressed genes and functional enrichment analysis in SKCM. (A) PCC was used to calculate correlations between CXCL13 and expressed genes in SKCM. Heat map showing top 30 genes positively correlated with CXCL13 in SKCM. (B) The significantly enriched GO annotations of the top 30 high CXCL13 co-expression genes in SKCM were analyzed. Figure 3A was produced using pheatmap packages in R (version 3.6, https://www.r-project.org/) and Figure 3B was produced using clusterProfiler packages in R (version 3.6, https://www.r-project.org/).
Figure 3. CXCL13 associated top 30 co-expressed genes and functional enrichment analysis in SKCM. (A) PCC was used to calculate correlations between CXCL13 and expressed genes in SKCM. Heat map showing top 30 genes positively correlated with CXCL13 in SKCM. (B) The significantly enriched GO annotations of the top 30 high CXCL13 co-expression genes in SKCM were analyzed. Figure 3A was produced using pheatmap packages in R (version 3.6, https://www.r-project.org/) and Figure 3B was produced using clusterProfiler packages in R (version 3.6, https://www.r-project.org/). 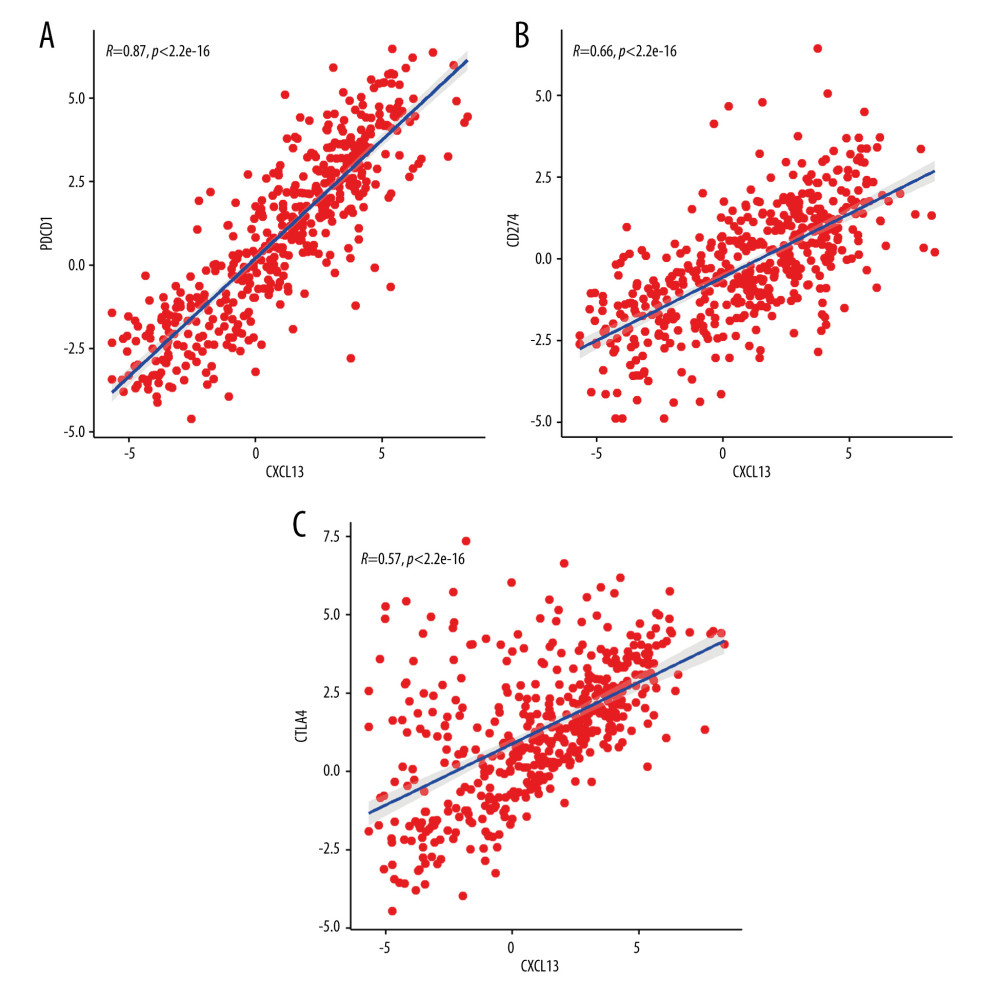 Figure 4. Gene expression correlation analysis. The scatter plot shows Pearson correlation of CXCL13 expression with expression of PDCD1 (A), CD274 (B), and CTLA4 (C). These figures were produced using ggplot2, ggpubr, and ggpmisc packages in R (version 3.6, https://www.r-project.org/).
Figure 4. Gene expression correlation analysis. The scatter plot shows Pearson correlation of CXCL13 expression with expression of PDCD1 (A), CD274 (B), and CTLA4 (C). These figures were produced using ggplot2, ggpubr, and ggpmisc packages in R (version 3.6, https://www.r-project.org/). 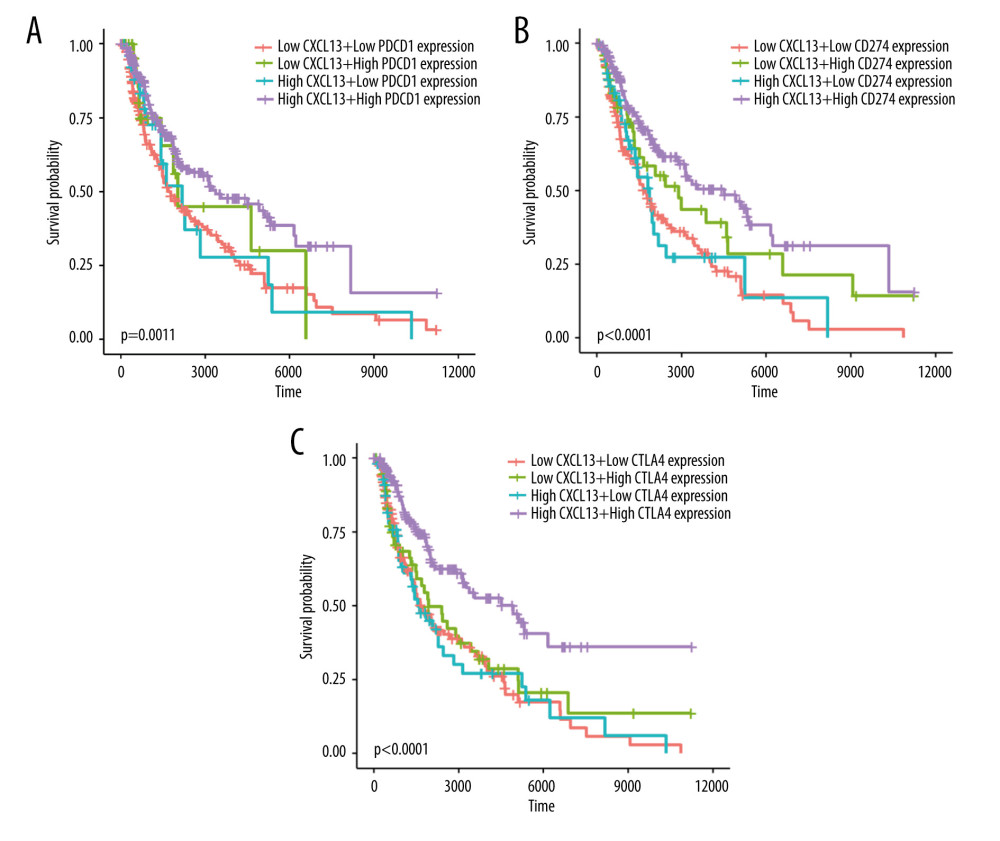 Figure 5. Kaplan-Meier survival curves show that CXCL13 expression and PDCD1/CD274/CTLA4 expression are significantly associated with OS in TCGA-SKCM cohorts (A) PDCD1, (B) CD274, (C) CTLA4. These figures were produced using survival and survminer packages in R (version 3.6, https://www.r-project.org/).
Figure 5. Kaplan-Meier survival curves show that CXCL13 expression and PDCD1/CD274/CTLA4 expression are significantly associated with OS in TCGA-SKCM cohorts (A) PDCD1, (B) CD274, (C) CTLA4. These figures were produced using survival and survminer packages in R (version 3.6, https://www.r-project.org/). 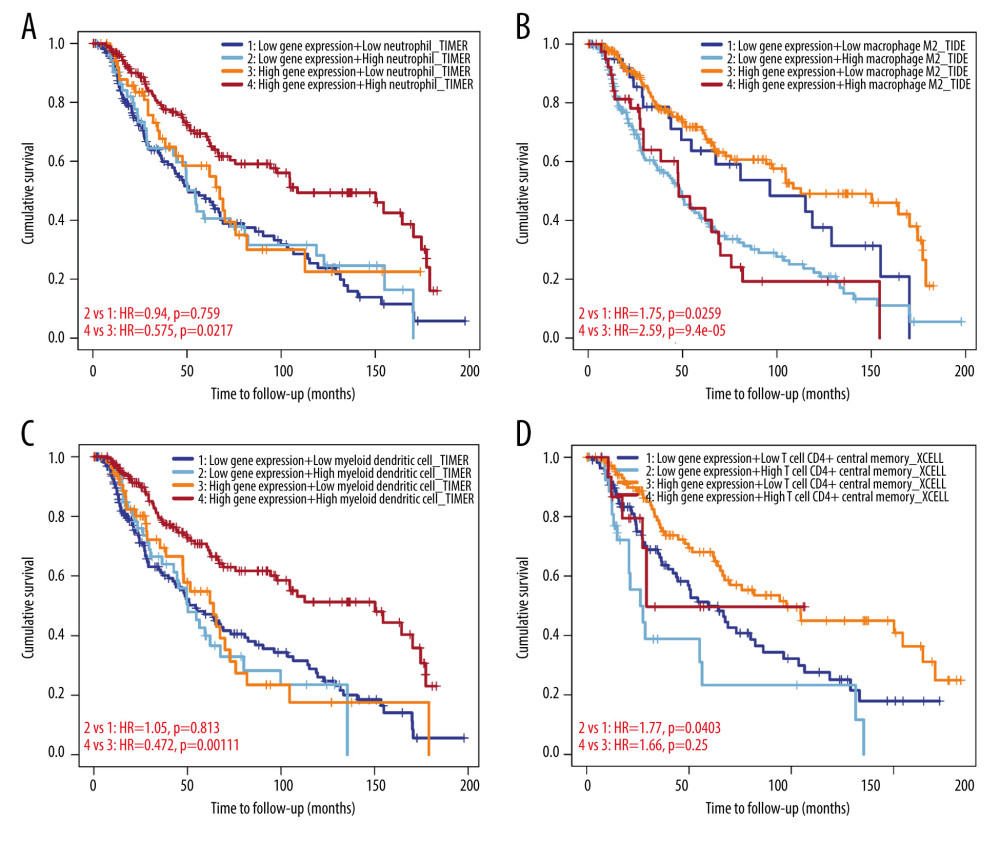 Figure 6. Correlation of CXCL13 expression with immune infiltration level in the TIMER database. (A) Neutrophil cells, (B) macrophages, (C) myeloid dendritic cells and (D) CD4+ T cells. These figures were produced using the TIMMER website (http://timer.cistrome.org, version 2.0).
Figure 6. Correlation of CXCL13 expression with immune infiltration level in the TIMER database. (A) Neutrophil cells, (B) macrophages, (C) myeloid dendritic cells and (D) CD4+ T cells. These figures were produced using the TIMMER website (http://timer.cistrome.org, version 2.0). References
1. Olszanski AJ, Current and future roles of targeted therapy and immunotherapy in advanced melanoma: J Manag Care Spec Pharm, 2014; 20; 346-56
2. Leonardi GC, Falzone L, Salemi R, Cutaneous melanoma: From pathogenesis to therapy (review): Int J Oncol, 2018; 52; 1071-80
3. Czarnecka AM, Bartnik E, Fiedorowicz M, Rutkowski P, Targeted therapy in melanoma and mechanisms of resistance: Int J Mol Sci, 2020; 21; 4576
4. Wolchok JD, Kluger H, Callahan MK, Nivolumab plus ipilimumab in advanced melanoma: N Engl J Med, 2013; 369; 122-33
5. Owen CN, Shoushtari AN, Chauhan D, Management of early melanoma recurrence despite adjuvant anti-PD-1 antibody therapy: Ann Oncol, 2020; 31; 1075-82
6. Chow MT, Luster AD, Chemokines in cancer: Cancer Immunol Res, 2014; 2; 1125-31
7. Keshava Prasad TS, Goel R, Kandasamy K, Human protein reference database – 2009 update: Nucleic Acids Res, 2009; 37; D767-72
8. Shannon P, Markiel A, Ozier O, Cytoscape: A software environment for integrated models of biomolecular interaction networks: Genome Res, 2003; 13; 2498-504
9. Li T, Fan J, Wang B, TIMER: A web server for comprehensive analysis of tumor-infiltrating immune cells: Cancer Res, 2017; 77; e108-10
10. Hussain M, Adah D, Tariq M, CXCL13/CXCR5 signaling axis in cancer: Life Sci, 2019; 227; 175-86
11. Huang B, Han W, Sheng ZF, Shen GL, Identification of immune-related biomarkers associated with tumorigenesis and prognosis in cutaneous melanoma patients: Cancer Cell Int, 2020; 20; 195
12. Zhou X, Peng M, He Y, CXC Chemokines as therapeutic targets and prognostic biomarkers in skin cutaneous melanoma microenvironment: Front Oncol, 2021; 11; 619003
13. Tauriello DVF, Palomo-Ponce S, Stork D, TGFβ drives immune evasion in genetically reconstituted colon cancer metastasis: Nature, 2018; 554; 538-43
14. Patel SP, Kurzrock R, PD-L1 expression as a predictive biomarker in cancer immunotherapy: Mol Cancer Ther, 2015; 14; 847-56
15. Jacquelot N, Duong CPM, Belz GT, Zitvogel L, Targeting chemokines and chemokine receptors in melanoma and other cancers: Front Immunol, 2018; 9; 2480
16. Kazanietz MG, Durando M, Cooke M, CXCL13 and its receptor CXCR5 in cancer: Inflammation, immune response, and beyond: Front Endocrinol (Lausanne), 2019; 10; 471
17. Razis E, Kalogeras KT, Kotsantis I, The role of CXCL13 and CXCL9 in early breast cancer: Clinical Breast Cancer, 2020; 20; e36-53
18. Huang B, Han W, Sheng Z-F, Shen G-L, Identification of immune-related biomarkers associated with tumorigenesis and prognosis in cutaneous melanoma patients: Cancer Cell Int, 2020; 20; 1-15
19. Jiao F, Sun H, Yang Q, Association of CXCL13 and immune cell infiltration signature in clear cell renal cell carcinoma: Int J Med Sci, 2020; 17; 1610-24
20. Rivadeneira DB, DePeaux K, Wang Y, Oncolytic viruses engineered to enforce leptin expression reprogram tumor-infiltrating T cell metabolism and promote tumor clearance: Immunity, 2019; 51; 548-60.e4
Figures
 Figure 1. Venn diagram and the most significant KEGG pathways of 41DEGs. (A) The Venn diagram displays the number of DEGs in 3 datasets from the GEO database. (B) Four significant KEGG pathways of 41 DEGs. Adjusted p value <0.01. Figure 1A was produced using the Venny website (https://bioinfogp.cnb.csic.es/tools/venny/index.html, version 2.1). Figure 1B was produced using clusterProfiler packages in R (version 3.6, https://www.r-project.org/).
Figure 1. Venn diagram and the most significant KEGG pathways of 41DEGs. (A) The Venn diagram displays the number of DEGs in 3 datasets from the GEO database. (B) Four significant KEGG pathways of 41 DEGs. Adjusted p value <0.01. Figure 1A was produced using the Venny website (https://bioinfogp.cnb.csic.es/tools/venny/index.html, version 2.1). Figure 1B was produced using clusterProfiler packages in R (version 3.6, https://www.r-project.org/). Figure 2. The network and survival plot. (A) The PPI (Protein-Protein Interaction) network of 4 high-degree DEGs (differentially-expressed genes). (B) Kaplan-Meier survival curves show that the expression of CXCL13 is consistently associated with better overall survival (OS) in SKCM. Figure 2A was produced using Cytoscape (version 3.8.2, https://cytoscape.org/). Figure 2B was produced using survival and survminer packages in R (version 3.6, https://www.r-project.org/).
Figure 2. The network and survival plot. (A) The PPI (Protein-Protein Interaction) network of 4 high-degree DEGs (differentially-expressed genes). (B) Kaplan-Meier survival curves show that the expression of CXCL13 is consistently associated with better overall survival (OS) in SKCM. Figure 2A was produced using Cytoscape (version 3.8.2, https://cytoscape.org/). Figure 2B was produced using survival and survminer packages in R (version 3.6, https://www.r-project.org/). Figure 3. CXCL13 associated top 30 co-expressed genes and functional enrichment analysis in SKCM. (A) PCC was used to calculate correlations between CXCL13 and expressed genes in SKCM. Heat map showing top 30 genes positively correlated with CXCL13 in SKCM. (B) The significantly enriched GO annotations of the top 30 high CXCL13 co-expression genes in SKCM were analyzed. Figure 3A was produced using pheatmap packages in R (version 3.6, https://www.r-project.org/) and Figure 3B was produced using clusterProfiler packages in R (version 3.6, https://www.r-project.org/).
Figure 3. CXCL13 associated top 30 co-expressed genes and functional enrichment analysis in SKCM. (A) PCC was used to calculate correlations between CXCL13 and expressed genes in SKCM. Heat map showing top 30 genes positively correlated with CXCL13 in SKCM. (B) The significantly enriched GO annotations of the top 30 high CXCL13 co-expression genes in SKCM were analyzed. Figure 3A was produced using pheatmap packages in R (version 3.6, https://www.r-project.org/) and Figure 3B was produced using clusterProfiler packages in R (version 3.6, https://www.r-project.org/). Figure 4. Gene expression correlation analysis. The scatter plot shows Pearson correlation of CXCL13 expression with expression of PDCD1 (A), CD274 (B), and CTLA4 (C). These figures were produced using ggplot2, ggpubr, and ggpmisc packages in R (version 3.6, https://www.r-project.org/).
Figure 4. Gene expression correlation analysis. The scatter plot shows Pearson correlation of CXCL13 expression with expression of PDCD1 (A), CD274 (B), and CTLA4 (C). These figures were produced using ggplot2, ggpubr, and ggpmisc packages in R (version 3.6, https://www.r-project.org/). Figure 5. Kaplan-Meier survival curves show that CXCL13 expression and PDCD1/CD274/CTLA4 expression are significantly associated with OS in TCGA-SKCM cohorts (A) PDCD1, (B) CD274, (C) CTLA4. These figures were produced using survival and survminer packages in R (version 3.6, https://www.r-project.org/).
Figure 5. Kaplan-Meier survival curves show that CXCL13 expression and PDCD1/CD274/CTLA4 expression are significantly associated with OS in TCGA-SKCM cohorts (A) PDCD1, (B) CD274, (C) CTLA4. These figures were produced using survival and survminer packages in R (version 3.6, https://www.r-project.org/). Figure 6. Correlation of CXCL13 expression with immune infiltration level in the TIMER database. (A) Neutrophil cells, (B) macrophages, (C) myeloid dendritic cells and (D) CD4+ T cells. These figures were produced using the TIMMER website (http://timer.cistrome.org, version 2.0).
Figure 6. Correlation of CXCL13 expression with immune infiltration level in the TIMER database. (A) Neutrophil cells, (B) macrophages, (C) myeloid dendritic cells and (D) CD4+ T cells. These figures were produced using the TIMMER website (http://timer.cistrome.org, version 2.0). In Press
07 Mar 2024 : Clinical Research
Knowledge of and Attitudes Toward Clinical Trials: A Questionnaire-Based Study of 179 Male Third- and Fourt...Med Sci Monit In Press; DOI: 10.12659/MSM.943468
08 Mar 2024 : Animal Research
Modification of Experimental Model of Necrotizing Enterocolitis (NEC) in Rat Pups by Single Exposure to Hyp...Med Sci Monit In Press; DOI: 10.12659/MSM.943443
18 Apr 2024 : Clinical Research
Comparative Analysis of Open and Closed Sphincterotomy for the Treatment of Chronic Anal Fissure: Safety an...Med Sci Monit In Press; DOI: 10.12659/MSM.944127
08 Mar 2024 : Laboratory Research
Evaluation of Retentive Strength of 50 Endodontically-Treated Single-Rooted Mandibular Second Premolars Res...Med Sci Monit In Press; DOI: 10.12659/MSM.944110
Most Viewed Current Articles
17 Jan 2024 : Review article
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
14 Dec 2022 : Clinical Research
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387
01 Jan 2022 : Editorial
Editorial: Current Status of Oral Antiviral Drug Treatments for SARS-CoV-2 Infection in Non-Hospitalized Pa...DOI :10.12659/MSM.935952
Med Sci Monit 2022; 28:e935952