24 February 2021: Clinical Research
Expression of Cell Division Cycle Protein 45 in Tissue Microarrays and the Gene by Bioinformatics Analysis in Human Hepatocellular Carcinoma and Patient Outcomes
Hui-Ping Lu1ABCDEFG, Xiu-Fang Du1BCDEF, Jian-Di Li1BCDEF, Su-Ning Huang2ADEF, Rong-Quan He3ABDEF, Hua-Yu Wu4ABDEF, Ming-Fen Li5BCDEF, Wei-Zi Wu6CDEF, Ji-Tian Chen6BCDE, Wei-Jia Mo1ABCDEFG, Gang Chen1ABCDEFG*DOI: 10.12659/MSM.928800
Med Sci Monit 2021; 27:e928800
Abstract
BACKGROUND: Hepatocellular carcinoma (HCC) causes a heavy disease burden worldwide. Cell division cycle 45 (Cdc45) and its encoding gene (CDC45) have been studied for a long time, but their expression patterns and roles in liver carcinogenesis and advanced HCC deterioration are still incompletely understood. This study integrated tissue microarray and bioinformatics analyses to explore the expression and clinical value of CDC45 and Cdc45 in HCC.
MATERIAL AND METHODS: In HCC, the expression and relationships with clinic-pathological parameters of CDC45 and Cdc45 were investigated by integrating the RNA-sequencing data, downloaded from The Cancer Genome Atlas and Oncomine databases, and tissue microarray with immunohistochemistry staining. Co-expressed genes and genetic alterations of CDC45 separately obtained from Oncomine and cBioPortal databases were identified to shed light on the potential mechanisms of CDC45 in HCC.
RESULTS: CDC45 and Cdc45 were both overexpressed in HCC tissues, and the CDC45 level progressively increased from stage I to III. The survival outcomes of the group with high CDC45 expression were significantly worse compared with the group with low expression. Amplification and deep deletion were 2 major significant alteration types in HCC patients, and the outcomes were worse in patients with altered versus unaltered CDC45. NUDT1, E2F1, CCNE2, MCM5, and CENPM were identified as the most significantly co-expressed genes.
CONCLUSIONS: CDC45 and Cdc45 were both upregulated in HCC, and increased expression levels and genetic alternations of CDC45 were correlated with worse prognosis in HCC patients. CDC45 may promote HCC by co-expressing with NUDT1, E2F1, CCNE2, MCM5, and CENPM.
Keywords: Carcinoma, Hepatocellular, Immunohistochemistry, Tissue Array Analysis, Biomarkers, Tumor, Cell Cycle Proteins, Computational Biology, Gene Expression Profiling, Liver Neoplasms, Sequence Analysis, RNA
Background
In the United States, liver cancer has increased in mortality and morbidity more than any other human cancer during the past decade [1]. In China, new cases of hepatocellular carcinoma (HCC) represent about 45% of the total HCC cases reported worldwide annually [2,3]. Liver cancer has long been recognized as a serious threat to human health [4]. HCC is the most common subtype of liver cancer, and it is well known for its high heterogeneity, aggressiveness, fatality rate, and incidence, as well as a lack of acceptable and efficient biomarkers to predict disease progression and guide its treatment. Thus, most HCC cases are diagnosed at an advanced stage, and the 5-year survival rate of HCC patients in both the United States and China is less than 20% [1,3].
Growing research on tumor signal transduction pathways has demonstrated that HCC progression results from the aberrant activation of several molecules in various signaling pathways controlling cell cycle progression or arrest, proliferation, differentiation, cell survival, and apoptosis [5]. Therefore, it is urgent and essential to research cancer genes to better understand their involvement in the underlying molecular mechanisms of tumorigenesis and cancer progression and to develop more effective diagnosis and treatment options for HCC.
Cancer is characterized by uncontrolled cell proliferation and DNA replication catalyzed by DNA helicases, and in-depth research on these characteristic molecular events is needed to probe their underlying causes and design novel therapeutic targets. The gene cell division cycle 45 (
However, agreement has not been reached on the expression and functions of
Material and Methods
Clinical Implications of CDC45 mRNA Expression Levels in HCC Tissues
CDC45 MRNA EXPRESSION LEVELS IN HCC TISSUES BASED ON RNA-SEQUENCING DATA: In this study, FireBrowse [15] (http://firebrowse.org/) and GEPIA (Gene Expression Profiling Interactive) [16] (http://gepia.cancer-pku.cn/) were used to analyze all CDC45 mRNA expression data. Both are based on the RNA-sequencing data from The Cancer Genome Atlas (TCGA), and GEPIA also contains the data from the Genotype-Tissue Expression (GTEx) project. To identify the expression level of CDC45 among different types of cancer, RNA-Seq by Expectation-Maximization (RSEM) (log2) was used, which was shown in FireBrowse. With regard to GEPIA and the data from GTEx, the TPM (transcripts per million) format for the relevant CDC45 mRNA expression levels were calculated. In total, 369 cases of HCC and 160 cases of non-HCC liver samples were examined. The clinicopathological information for the HCC patients was downloaded from TCGA, including their clinical stages, overall survival rates, and disease-free survival numbers.
CDC45 MRNA EXPRESSION LEVELS IN HCC TISSUES BASED ON OTHER HIGH-THROUGHPUT DATASETS: The Oncomine (https://www.oncomine.org) database contains gene expression measurement from more than 4700 experiments and is a powerful public platform for oncogenomic analyses [17]. Therefore, to validate the CDC45 mRNA expression levels in HCC patients, we searched the Oncomine datasets for the CDC45 expression levels in HCC and non-HCC liver tissues. All microarray platforms and RNA-sequencing data were considered.
ANALYZING CDC45 EXPRESSION IN HCC WITH IN-HOUSE TISSUE MICROARRAYS AND IHC:
IHC was performed according to the manufacturer’s instructions. All patients had given informed consent, and this study was authorized by the Ethics Committee of the First Affiliated Hospital of Guangxi Medical University in Nanning, China (No. 2020 [KY-E-126]). The formalin-fixed paraffin-embedded tissue microarray (TMA) was supplied by Fanpu Biotech, Inc. (LVC1021, LVC1601, Guilin, China) and contained 137 HCC tissues and 62 non-HCC liver tissues. After being deparaffinized and rehydrated, the tissue slides were put into a boiled 0.01 M citrate buffer (pH 6.0) to retrieve the antigen. The endogenous peroxidase activity was blocked with 3% H2O2. The primary antibody against Cdc45 was a rabbit anti-human Cdc45 monoclonal antibody (ab126762) (Abcam, UK, dilution 1: 100), which was incubated overnight at 4°C and then with horseradish peroxidase (HRP)-labeled secondary antibody (ready-to-use, Long Island Antibody, Shanghai, China) at room temperature for 25 min. The last step involved the visualization of the HRP with 3,3′-diaminobenzidine (DAB), followed by dehydration, sealing, and evaluation with a bright-field microscope. Brown-yellow granules in the nucleus and/or cytoplasm indicated positive staining. Negative control sections were incubated with phosphate-buffered saline during the primary antibody incubation.
The 2 stained TMAs were assessed blindly by 2 independent pathologists and were each evaluated at ×400 magnification for the intensity and percentage of positive cells in 10 randomly selected fields. The staining intensity was scored according to a model used in prior studies [18], with the recorded percentage falling into 1 of 4 grades: 0 (<5%), 1 (5–25%), 2 (26–50%), or 3 (>50%). To find the total IHC score, the intensity score was multiplied by the percentage score.
THE CO-EXPRESSED GENES OF CDC45 IN HCC TISSUES: Since CDC45 may be co-expressed with other genes and mediate a potential mechanism in the pathogenesis and progression of HCC, we investigated the genes co-expressed with CDC45 in 102 cases of HCC from the Chen Liver dataset, 35 cases of HCC from the Wurmbach Liver dataset, and 22 cases of HCC from the Roessler Liver dataset. Heat-maps were drawn to show the most significant co-expressed genes based on their correlation coefficient. All these analyses were based on Oncomine [17].
THE GENETIC ALTERATIONS OF CDC45 IN HCC TISSUES: Genetic alterations also contribute to the function of a gene. Thus, we used multiple datasets to reveal the genetic alterations of CDC45 in HCC. The following datasets from cBioPortal (The cBio Cancer Genomics Portal) [19] (http://cbioportal.org) were included: “Hepatocellular Carcinoma” (MSK, Clin Cancer Res 2018, 127 samples), “Hepatocellular Carcinoma” (INSERM, Nat Genet 2015, 243 samples), “Liver Hepatocellular Adenoma and Carcinomas” (MSK, PLoS One 2018, 19 samples), “Liver Hepatocellular Carcinoma” (AMC, Hepatology 2014, 231 samples), “Liver Hepatocellular Carcinoma” (RIKEN, Nat Genet 2012, 27 samples), and “Liver Hepatocellular Carcinoma” (TCGA, Firehose Legacy, 442 samples). The molecular profiles of the mutations, copy number of the alterations, and mRNA Expression z-Scores (RNA-Seq V2 RSEM) were analyzed [20].
STATISTICAL ANALYSIS:
All statistics were analyzed with SPSS Statistics (version 23.0). The results were presented as mean±standard deviation (SD). A
Results
The Clinical Significance of CDC45 mRNA Expression Levels in HCC
CALCULATING THE RNA-SEQUENCING DATA TO IDENTIFY THE CDC45 MRNA EXPRESSION LEVELS IN HCC TISSUES: Using the TCGA RNA-Seq data, FireBrowse provided the CDC45 mRNA expression levels for more than 30 types of tumors. In 27 types of tumors with nontumorous controls, the CDC45 mRNA expression levels were clearly upregulated compared with the controls (Figure 1A). To improve the accuracy and validity of the statistical analysis, we increased the number of noncancerous cases used in the comparison by adding normal liver samples from GTEx in GEPIA. The CDC45 mRNA expression levels were predominantly higher in the 369 cases of HCC than in the 160 cases of non-HCC liver controls (Figure 1B). We also found increases in the CDC45 mRNA expression levels from stage I to stage III, indicating that CDC45 might be responsible for tumor progression. However, at stage IV, the CDC45 mRNA expression levels decreased, probably due to the small sample size (Figure 1C). The proposed role of CDC45 mRNA in tumor development was also supported by the prognostic values obtained. The HR was 1.9 (P=0.00039) for the overall survival prediction (Figure 1D) and 1.6 for the disease-free survival prediction (P=0.0037, Figure 1E). The RNA-sequencing data suggested that the upregulation of CDC45 mRNA expression levels might play a part in causing tumorigenesis and poorer prognosis of HCC patients.
MINING THE MICROARRAY DATA TO EVALUATE THE CDC45 MRNA EXPRESSION LEVELS IN HCC TISSUES: To gather more evidence supporting the proposed oncogenic implications of CDC45 mRNA in HCC, we collected several microarray datasets from Oncomine. Four of these datasets showed that the CDC45 mRNA expression levels were indeed obviously upregulated in the HCC samples compared with the non-HCC controls, including the Chen Liver, Wurmbach Liver, Roessler Liver, and Roessler Liver 2 datasets (Figure 2).
ASSESSING THE CDC45 EXPRESSION LEVELS IN HCC TISSUES WITH IN-HOUSE TISSUE MICROARRAYS AND IHC: We further validated the CDC45 mRNA expression levels by assessing their protein levels with in-house tissue microarrays and IHC. The positive signaling of the Cdc45 was located in the nucleus and cytoplasm. In line with the mRNA levels, the Cdc45 levels in the HCC tissues were also notably increased in comparison to those in the non-HCC tissues (4.635±2.051 vs 2.565±1.410, respectively; P=4.317E-14; Figure 3).
THE GENES CO-EXPRESSED WITH CDC45 IN HCC TISSUES: The genes co-expressed with CDC45 could partially reveal the potential functional mechanism of CDC45 in HCC. Two most significant genes co-expressed with CDC45 in the Chen Liver dataset were NUDT1 (r=0.713) and E2F1 (r=0.675). In addition, 16 other genes had the same correlation index (r=0.652), including NUF2, NEK2, and ARL4A (Figure 4). The first co-expressed gene listed in the Wurmbach Liver dataset was CCNE2, followed by 16 genes with the same r value (0.666), including NUP155, CKS1B, and CNIH4 (Figure 5). The first 2 co-expressed genes in the Roessler Liver dataset were MCM5 and CENPM (r=0.823), followed by 5 other genes with the same r value (0.779): GINS2, ASF1B, E2F8, RNASEH2A, and CDKN2C (Figure 6).
THE GENETIC ALTERATIONS OF CDC45 IN HCC TISSUES: Genetic alterations, such as mutations and amplifications, could contribute to the potential molecular mechanism of CDC45 in HCC. To investigate this idea, we examined the genetic alterations of CDC45 in multiple datasets. CDC45 was found to be altered in 34 of 360 patients/complete samples (9%) included in 442 HCC cases from the TCGA dataset (Figure 7A). The patterns of CDC45 genetic alterations covered missense mutations, truncating mutations, amplification, deep deletion, and higher mRNA expression. Of note, the cases involving CDC45 genetic alterations tended to have more unfavorable overall survival (Figure 7B) and disease-free survival (Figure 7C). In addition to the TCGA datasets, we used 5 other studies to obtain CDC45 genetic alteration information (Figure 8A). CDC45 mutations could also be noted in liver (AMC) and HCC (Inserm, 2015) studies (Figure 8B). Across these datasets of CDC45 genetic alterations, a poorer overall survival rate was observed in the cases involving CDC45 alterations than in the unaltered group (Figure 9A), although a poorer disease-free survival rate was not observed (Figure 9B).
Discussion
Overexpressed
The role of
The mechanism of
There were 2 main limitations to this study. First, the roles of
Conclusions
The present study confirmed the upregulated expression pattern of
Figures
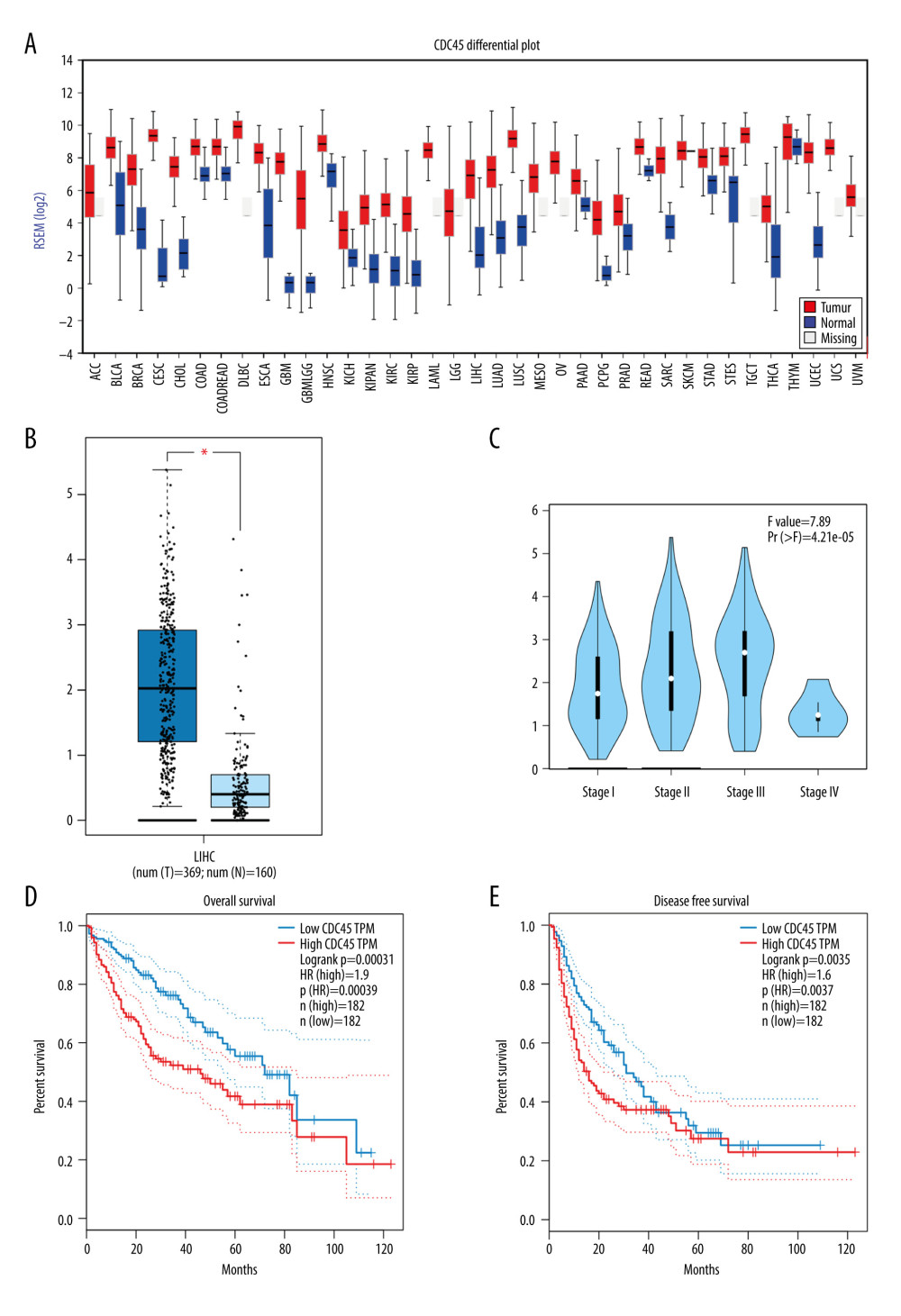 Figure 1. The clinical significance of CDC45 mRNA expression levels in hepatocellular carcinoma (HCC) tissues based on RNA-Seq data. The mRNA expression level of CDC45 was clearly upregulated compared with the controls (A, B). (A) The CDC45 mRNA expression was clearly higher in HCC than in normal tissues. (B) The CDC45 mRNA expression was significantly increased in HCC compared with normal tissues. (C) The CDC45 mRNA expression levels progressively increased from stage I to stage III and decreased in stage IV. A high CDC45 mRNA expression level indicated a negative prognosis (D: for overall survival, HR=1.9, P=0.00039; E: for disease-free survival, HR=1.6, P=0.0037). HR=hazard ratio.
Figure 1. The clinical significance of CDC45 mRNA expression levels in hepatocellular carcinoma (HCC) tissues based on RNA-Seq data. The mRNA expression level of CDC45 was clearly upregulated compared with the controls (A, B). (A) The CDC45 mRNA expression was clearly higher in HCC than in normal tissues. (B) The CDC45 mRNA expression was significantly increased in HCC compared with normal tissues. (C) The CDC45 mRNA expression levels progressively increased from stage I to stage III and decreased in stage IV. A high CDC45 mRNA expression level indicated a negative prognosis (D: for overall survival, HR=1.9, P=0.00039; E: for disease-free survival, HR=1.6, P=0.0037). HR=hazard ratio. 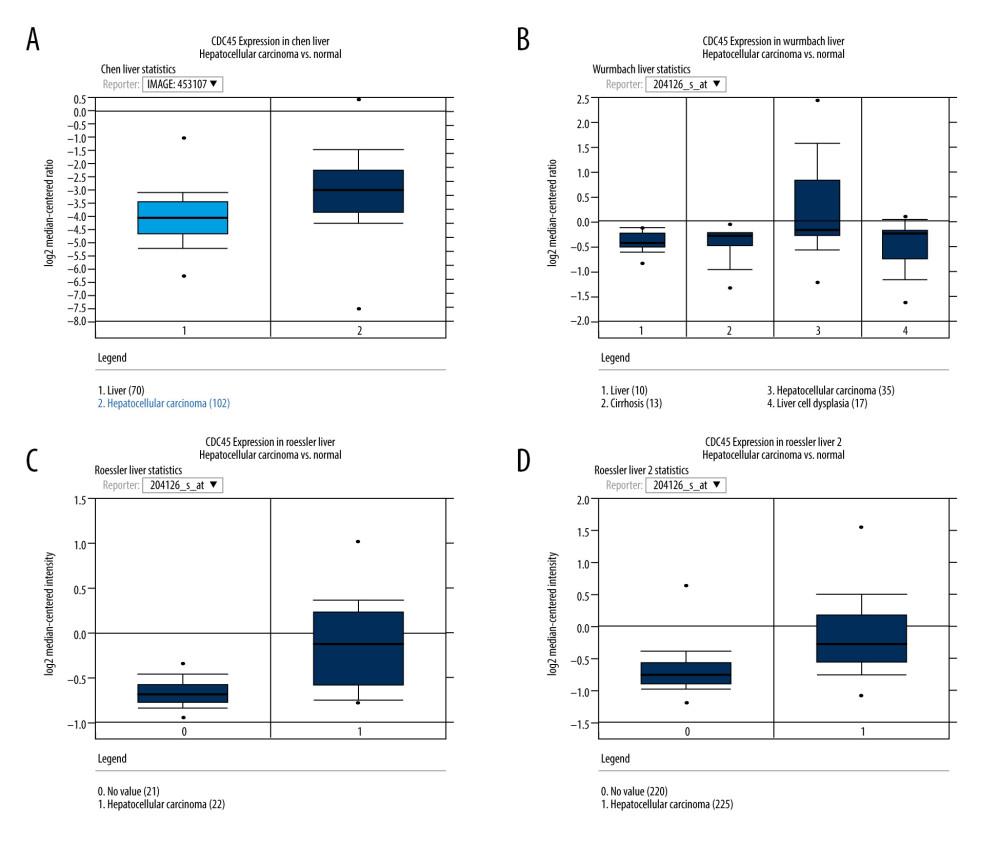 Figure 2. The CDC45 mRNA expression levels were obviously upregulated in the microarrays from the Oncomine datasets. (A) Chen Liver: fold change=2.135; P=1.16E-10; 197 samples; 10 802 measured genes; platform not predefined in Oncomine. (B) Wurmbach Liver: fold change=1.581; P=4.17E-5; 75 samples; 19 574 measured genes; Human Genome U133 Plus 2.0 Array. (C) Roessler Liver: fold change=1.429; P=2.70E-5; 43 samples; 12 603 measured genes; Human Genome U133A 2.0 Array. (D) Roessler Liver 2: fold change=1.453; P=2.00E-36; 445 samples; 12 624 measured genes; Affymetrix Human Genome HT U133A Array.
Figure 2. The CDC45 mRNA expression levels were obviously upregulated in the microarrays from the Oncomine datasets. (A) Chen Liver: fold change=2.135; P=1.16E-10; 197 samples; 10 802 measured genes; platform not predefined in Oncomine. (B) Wurmbach Liver: fold change=1.581; P=4.17E-5; 75 samples; 19 574 measured genes; Human Genome U133 Plus 2.0 Array. (C) Roessler Liver: fold change=1.429; P=2.70E-5; 43 samples; 12 603 measured genes; Human Genome U133A 2.0 Array. (D) Roessler Liver 2: fold change=1.453; P=2.00E-36; 445 samples; 12 624 measured genes; Affymetrix Human Genome HT U133A Array. 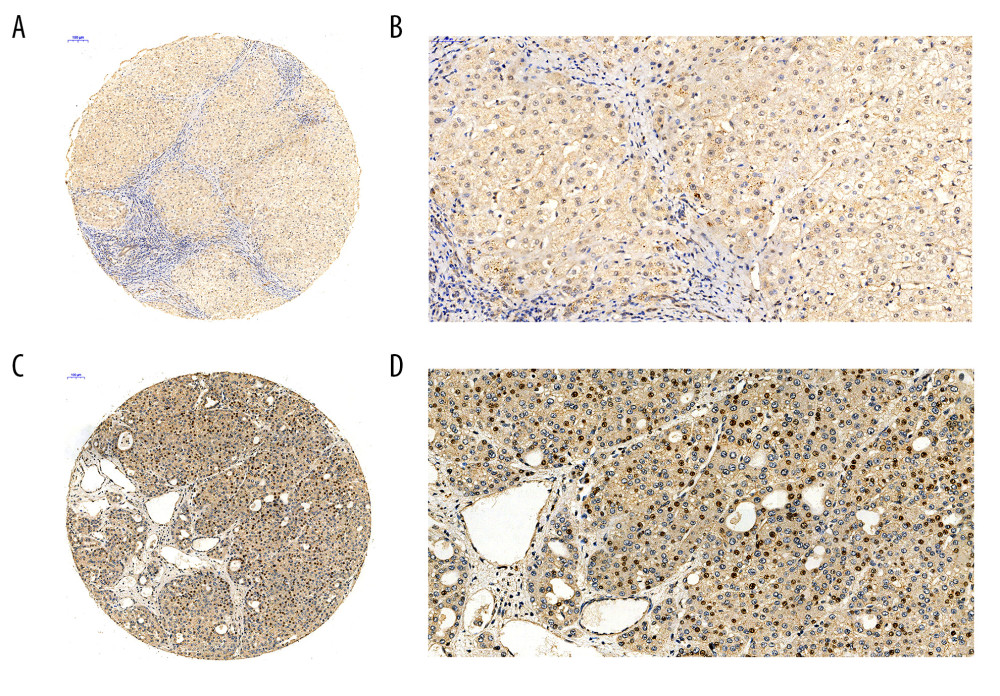 Figure 3. The Cdc45 expression levels for the in-house tissue microarrays. Immunohistochemistry staining to evaluate the expression of Cdc45 in hepatocellular carcinoma (HCC) and adjacent tissues. (A, B) The expression of Cdc45 was absent or weak in the non-HCC tissue (2.565±1.410). (C, D) The expression of Cdc45 was obviously high in the HCC tissue (4.635±2.051). Cdc45 was detected in the nucleus and cytoplasm. Magnification, ×20 (A, C) and ×400 (B, D).
Figure 3. The Cdc45 expression levels for the in-house tissue microarrays. Immunohistochemistry staining to evaluate the expression of Cdc45 in hepatocellular carcinoma (HCC) and adjacent tissues. (A, B) The expression of Cdc45 was absent or weak in the non-HCC tissue (2.565±1.410). (C, D) The expression of Cdc45 was obviously high in the HCC tissue (4.635±2.051). Cdc45 was detected in the nucleus and cytoplasm. Magnification, ×20 (A, C) and ×400 (B, D). 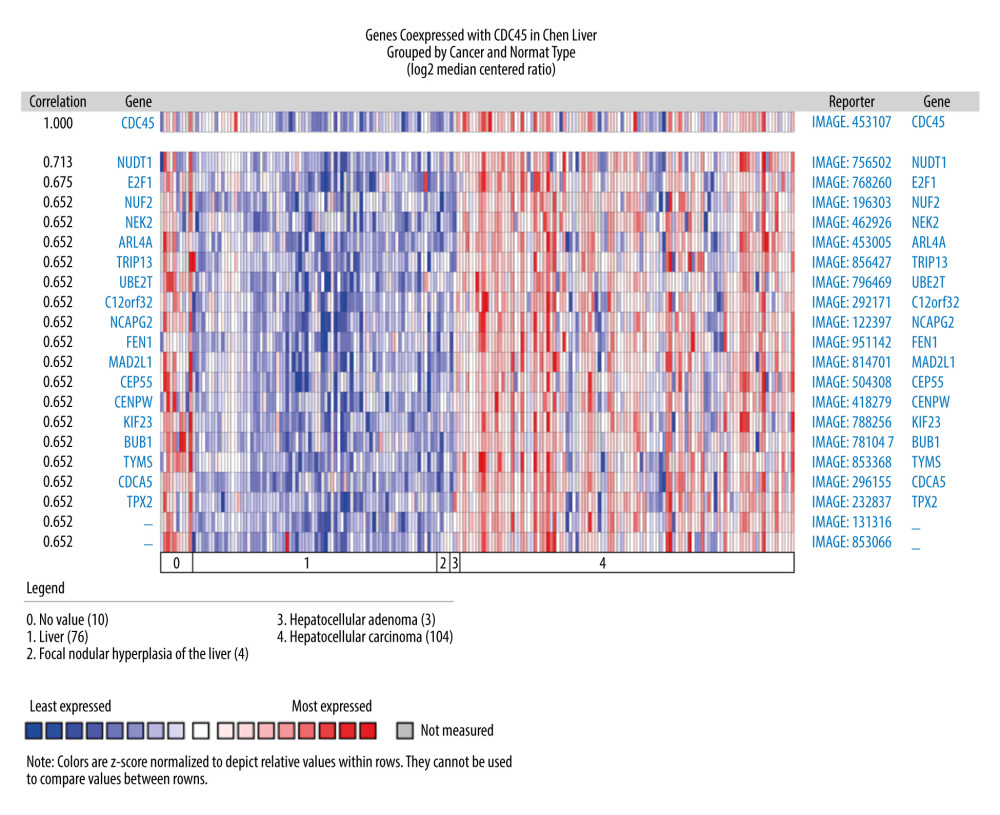 Figure 4. The co-expressed genes based on the Chen Liver microarray. NUDT1 and E2F1 were the top 2 genes on the list of co-expressed genes of CDC45 (r=0.713 and 0.675, respectively). The correlation index of other genes was 0.652.
Figure 4. The co-expressed genes based on the Chen Liver microarray. NUDT1 and E2F1 were the top 2 genes on the list of co-expressed genes of CDC45 (r=0.713 and 0.675, respectively). The correlation index of other genes was 0.652. 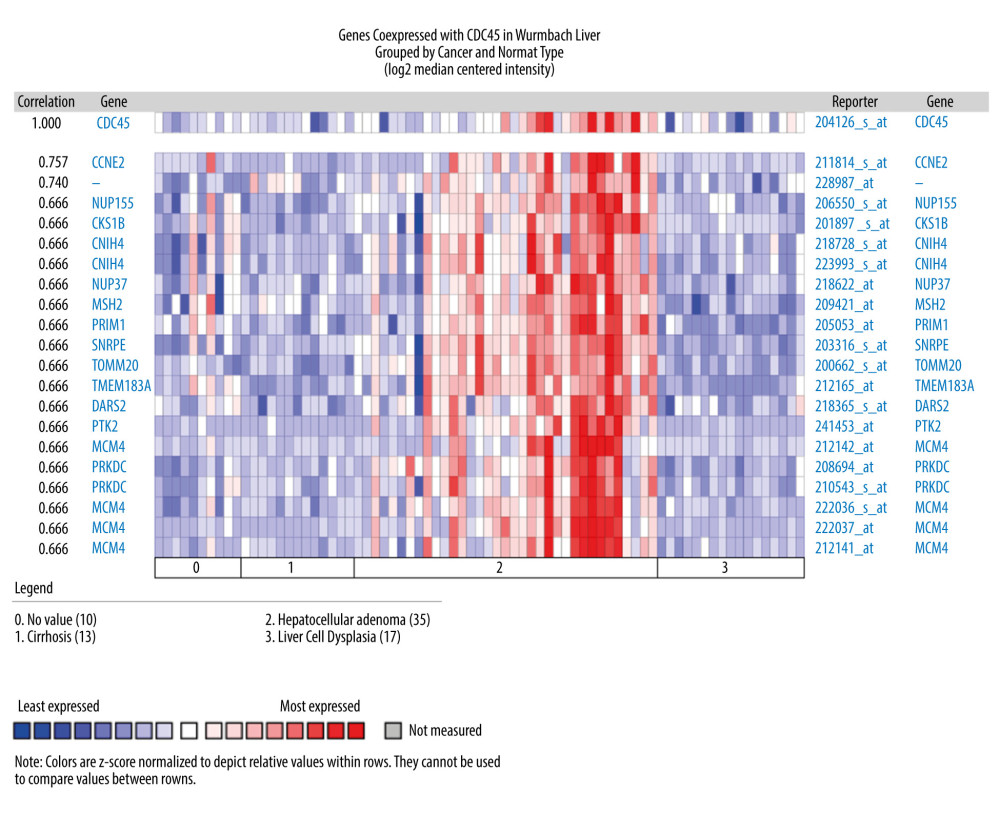 Figure 5. The co-expressed genes based on the Wurmbach Liver microarray. All the correlation indexes of co-expressed genes of CDC45 were 0.666, except CCNE2 (r=0.757).
Figure 5. The co-expressed genes based on the Wurmbach Liver microarray. All the correlation indexes of co-expressed genes of CDC45 were 0.666, except CCNE2 (r=0.757). 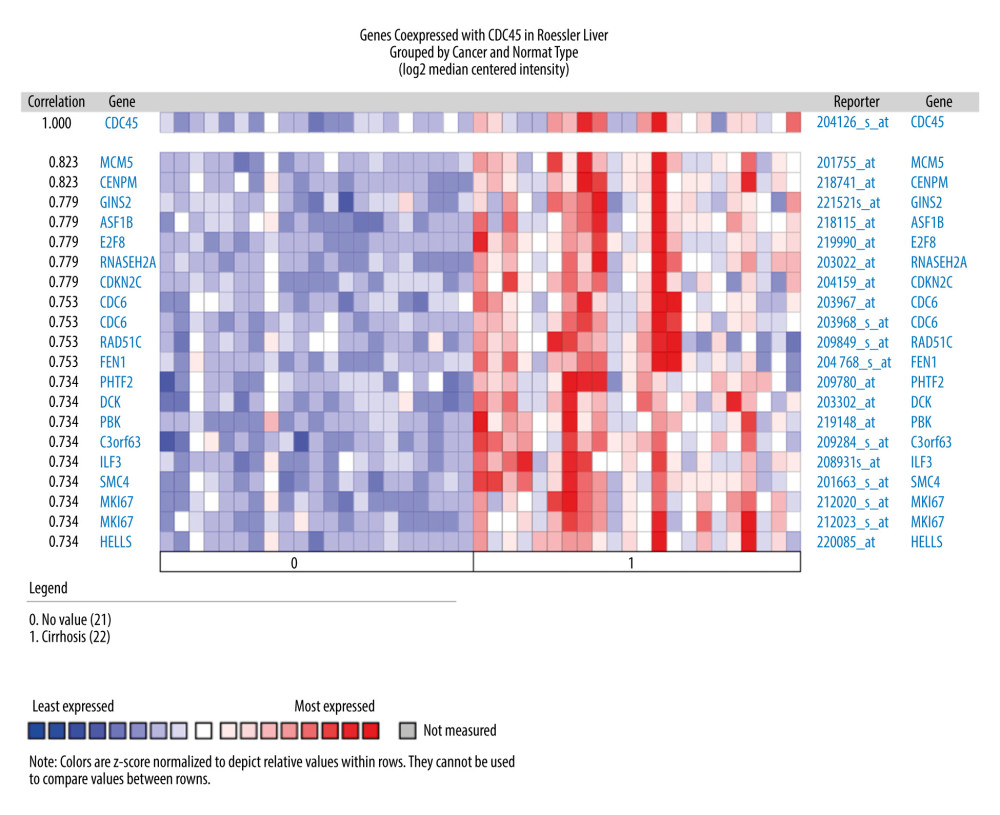 Figure 6. The co-expressed genes based on the Roessler Liver microarray. MCM5, CENPM, GINS2, ASF1B, E2F8, RNASEH2A, and CDKN2C were the top genes co-expressed with CDC45 in Roessler Liver.
Figure 6. The co-expressed genes based on the Roessler Liver microarray. MCM5, CENPM, GINS2, ASF1B, E2F8, RNASEH2A, and CDKN2C were the top genes co-expressed with CDC45 in Roessler Liver. 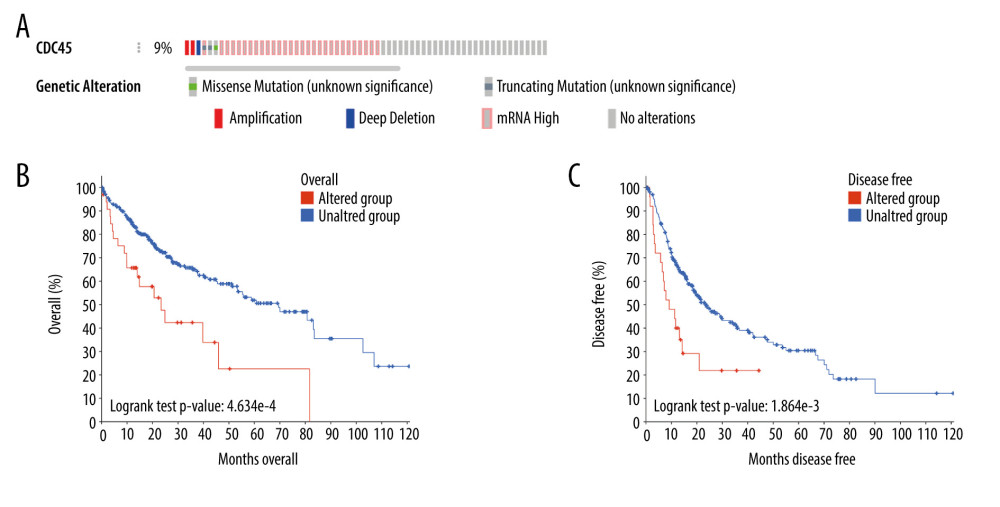 Figure 7. The clinical implications of CDC45 genetic alterations. (A) The oncoprint of CDC45 in 360 patients/complete samples from 442 cases of hepatocellular carcinoma (HCC). CDC45 was altered in 34 (9%) cases. Patients with the CDC45 genetic alteration were more likely to have poor lifespan (B, for overall survival; C, for disease-free survival).
Figure 7. The clinical implications of CDC45 genetic alterations. (A) The oncoprint of CDC45 in 360 patients/complete samples from 442 cases of hepatocellular carcinoma (HCC). CDC45 was altered in 34 (9%) cases. Patients with the CDC45 genetic alteration were more likely to have poor lifespan (B, for overall survival; C, for disease-free survival). 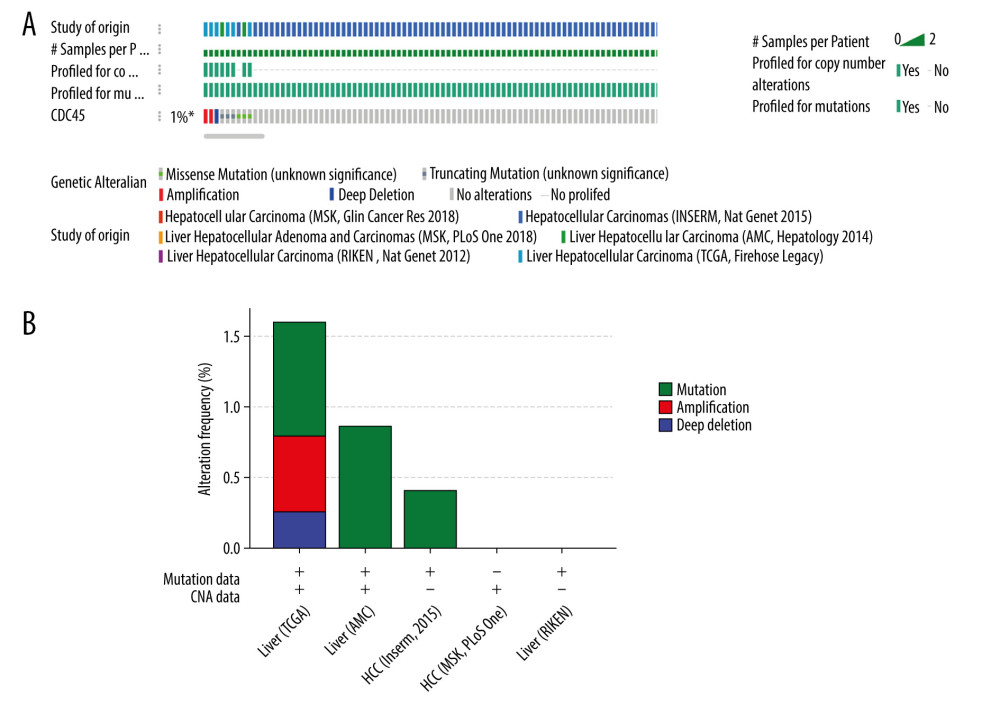 Figure 8. The CDC45 genetic alterations in multiple HCC datasets. (A) The oncoprint of CDC45 in 1089 cases of hepatocellular carcinoma (HCC) from multiple datasets involved The Cancer Genome Atlas (TCGA), AMC, Inserm, MSK, and RIKEN. CDC45 was altered in 9 (1%) of the patients. (B) The alteration frequency of CDC45 in HCC and mutation was the most common alternate forms, especially in liver (AMC) and HCC (Inserm, 2015) studies.
Figure 8. The CDC45 genetic alterations in multiple HCC datasets. (A) The oncoprint of CDC45 in 1089 cases of hepatocellular carcinoma (HCC) from multiple datasets involved The Cancer Genome Atlas (TCGA), AMC, Inserm, MSK, and RIKEN. CDC45 was altered in 9 (1%) of the patients. (B) The alteration frequency of CDC45 in HCC and mutation was the most common alternate forms, especially in liver (AMC) and HCC (Inserm, 2015) studies. 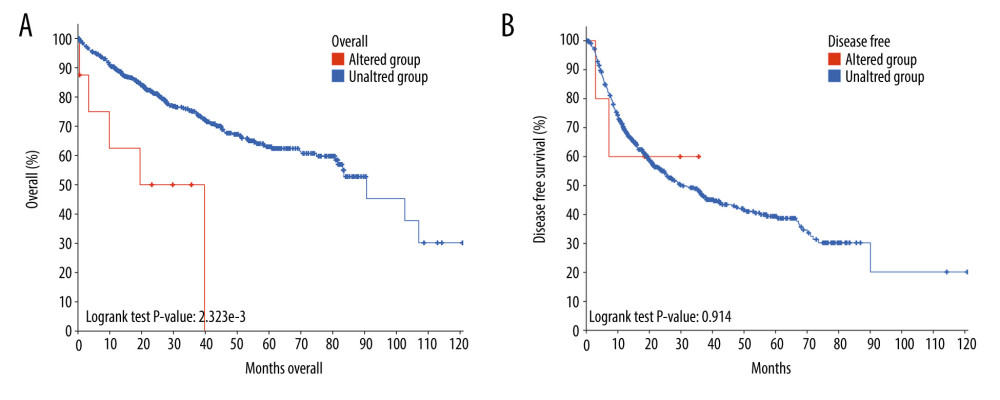 Figure 9. The prognostic values of CDC45 genetic alterations in hepatocellular carcinoma (HCC). The survival time of altered group was less than the unaltered group, both for overall survival (A) and disease-free survival (B).
Figure 9. The prognostic values of CDC45 genetic alterations in hepatocellular carcinoma (HCC). The survival time of altered group was less than the unaltered group, both for overall survival (A) and disease-free survival (B). References
1. Siegel RL, Miller KD, Jemal A, Cancer statistics, 2020: Cancer J Clin, 2020; 70(1); 7-30
2. Chen W, Zheng R, Baade PD, Cancer statistics in China, 2015: Cancer J Clin, 2016; 66(2); 115-32
3. Li Z, Wang Y, Duan S, Expression of TBX3 in hepatocellular carcinoma and its clinical implication: Med Sci Monit, 2018; 24; 9324-33
4. Endeshaw M, Hallowell BD, Razzaghi H, Trends in liver cancer mortality in the United States: Dual burden among foreign- and US-born persons: Cancer, 2019; 125(5); 726-34
5. Chen C, Wang G, Mechanisms of hepatocellular carcinoma and challenges and opportunities for molecular targeted therapy: World J Hepatol, 2015; 7(15); 1964-70
6. Seo YS, Kang YH, The human replicative helicase, the CMG complex, as a target for anti-cancer therapy: Front Mol Biosci, 2018; 5; 26
7. Broderick R, Nasheuer HP, Regulation of Cdc45 in the cell cycle and after DNA damage: Biochem Soc Trans, 2009; 37(Pt 4); 926-30
8. Pollok S, Bauerschmidt C, Sanger J, Human Cdc45 is a proliferation-associated antigen: FEBS J, 2007; 274(14); 3669-84
9. Nepon-Sixt BS, Bryant VL, Alexandrow MG, Myc-driven chromatin accessibility regulates Cdc45 assembly into CMG helicases: Commun Biol, 2019; 2; 110
10. Srinivasan SV, Dominguez-Sola D, Wang LC, Cdc45 is a critical effector of myc-dependent DNA replication stress: Cell Rep, 2013; 3(5); 1629-39
11. Tanaka S, Nakato R, Katou Y, Origin association of Sld3, Sld7, and Cdc45 proteins is a key step for determination of origin-firing timing: Curr Biol, 2011; 21(24); 2055-63
12. Wang Z, Yang B, Zhang M, lncRNA epigenetic landscape analysis identifies EPIC1 as an oncogenic lncRNA that interacts with MYC and promotes cell-cycle progression in cancer: Cancer Cell, 2018; 33(4); 706-20e9
13. Xiang XH, Yang L, Zhang X, Seven-senescence-associated gene signature predicts overall survival for Asian patients with hepatocellular carcinoma: World J Gastroenterol, 2019; 25(14); 1715-28
14. Yang C, Xie S, Wu Y, Prognostic implications of CDC45 expression in hepatocellular carcinoma: Research Square;, 2020 https://www.researchsquare.com/article/rs-31516/v1
15. Deng M, Bragelmann J, Kryukov I, FirebrowseR: An R client to the Broad Institute’s Firehose Pipeline: Database (Oxford), 2017; 2017; baw160
16. Tang Z, Li C, Kang B, GEPIA. A web server for cancer and normal gene expression profiling and interactive analyses: Nucleic Acids Res, 2017; 45(W1); W98-102
17. Rhodes DR, Kalyana-Sundaram S, Mahavisno V, Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles: Neoplasia, 2007; 9(2); 166-80
18. Luo B, Huang L, Gu Y, Expression of exportin-1 in diffuse large B-cell lymphoma: Immunohistochemistry and TCGA analyses: Int J Clin Exp Pathol, 2018; 11(12); 5547-60
19. Cerami E, Gao J, Dogrusoz U, The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data: Cancer Discov, 2012; 2(5); 401-4
20. Gao J, Aksoy BA, Dogrusoz U, Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal: Sci Signal, 2013; 6(269); pl1
21. Yang S, Ren X, Liang Y, KNK437 restricts the growth and metastasis of colorectal cancer via targeting DNAJA1/CDC45 axis: Oncogene, 2020; 39(2); 249-61
22. Sun J, Shi R, Zhao S, Cell division cycle 45 promotes papillary thyroid cancer progression via regulating cell cycle: Tumour Biol, 2017; 39(5) 1010428317705342
23. Li JN, Feng CJ, Lu YJ, mRNA expression of the DNA replication-initiation proteins in epithelial dysplasia and squamous cell carcinoma of the tongue: BMC Cancer, 2008; 8; 395
24. Hu Y, Wang L, Li Z, Potential prognostic and diagnostic values of CDC6, CDC45, ORC6 and SNHG7 in colorectal cancer: Onco Targets Ther, 2019; 12; 11609-21
25. Li HY, Jin N, Han YP, Jin XF, Pathway crosstalk analysis in prostate cancer based on protein-protein network data: Neoplasma, 2017; 64(1); 22-31
26. Gu Y, Wang X, Liu H, SET7/9 promotes hepatocellular carcinoma progression through regulation of E2F1: Oncol Rep, 2018; 40(4); 1863-74
27. Ou Q, Ma N, Yu Z, Nudix hydrolase 1 is a prognostic biomarker in hepatocellular carcinoma: Aging (Albany NY), 2020; 12(8); 7363-79
28. Yu Z, Wang R, Chen F, Five novel oncogenic signatures could be utilized as AFP-related diagnostic biomarkers for hepatocellular carcinoma based on next-generation sequencing: Dig Dis Sci, 2018; 63(4); 945-57
29. Sonntag R, Giebeler N, Nevzorova YA, Cyclin E1 and cyclin-dependent kinase 2 are critical for initiation, but not for progression of hepatocellular carcinoma: Proc Natl Acad Sci USA, 2018; 115(37); 9282-87
30. Xiao Y, Najeeb RM, Ma D, Upregulation of CENPM promotes hepatocarcinogenesis through mutiple mechanisms: J Exp Clin Cancer Res, 2019; 38(1); 458
31. Sun Q, Yu R, Wang C, Circular RNA circ-CSPP1 regulates CCNE2 to facilitate hepatocellular carcinoma cell growth via sponging miR-577: Cancer Cell Int, 2020; 20; 202
Figures
 Figure 1. The clinical significance of CDC45 mRNA expression levels in hepatocellular carcinoma (HCC) tissues based on RNA-Seq data. The mRNA expression level of CDC45 was clearly upregulated compared with the controls (A, B). (A) The CDC45 mRNA expression was clearly higher in HCC than in normal tissues. (B) The CDC45 mRNA expression was significantly increased in HCC compared with normal tissues. (C) The CDC45 mRNA expression levels progressively increased from stage I to stage III and decreased in stage IV. A high CDC45 mRNA expression level indicated a negative prognosis (D: for overall survival, HR=1.9, P=0.00039; E: for disease-free survival, HR=1.6, P=0.0037). HR=hazard ratio.
Figure 1. The clinical significance of CDC45 mRNA expression levels in hepatocellular carcinoma (HCC) tissues based on RNA-Seq data. The mRNA expression level of CDC45 was clearly upregulated compared with the controls (A, B). (A) The CDC45 mRNA expression was clearly higher in HCC than in normal tissues. (B) The CDC45 mRNA expression was significantly increased in HCC compared with normal tissues. (C) The CDC45 mRNA expression levels progressively increased from stage I to stage III and decreased in stage IV. A high CDC45 mRNA expression level indicated a negative prognosis (D: for overall survival, HR=1.9, P=0.00039; E: for disease-free survival, HR=1.6, P=0.0037). HR=hazard ratio. Figure 2. The CDC45 mRNA expression levels were obviously upregulated in the microarrays from the Oncomine datasets. (A) Chen Liver: fold change=2.135; P=1.16E-10; 197 samples; 10 802 measured genes; platform not predefined in Oncomine. (B) Wurmbach Liver: fold change=1.581; P=4.17E-5; 75 samples; 19 574 measured genes; Human Genome U133 Plus 2.0 Array. (C) Roessler Liver: fold change=1.429; P=2.70E-5; 43 samples; 12 603 measured genes; Human Genome U133A 2.0 Array. (D) Roessler Liver 2: fold change=1.453; P=2.00E-36; 445 samples; 12 624 measured genes; Affymetrix Human Genome HT U133A Array.
Figure 2. The CDC45 mRNA expression levels were obviously upregulated in the microarrays from the Oncomine datasets. (A) Chen Liver: fold change=2.135; P=1.16E-10; 197 samples; 10 802 measured genes; platform not predefined in Oncomine. (B) Wurmbach Liver: fold change=1.581; P=4.17E-5; 75 samples; 19 574 measured genes; Human Genome U133 Plus 2.0 Array. (C) Roessler Liver: fold change=1.429; P=2.70E-5; 43 samples; 12 603 measured genes; Human Genome U133A 2.0 Array. (D) Roessler Liver 2: fold change=1.453; P=2.00E-36; 445 samples; 12 624 measured genes; Affymetrix Human Genome HT U133A Array. Figure 3. The Cdc45 expression levels for the in-house tissue microarrays. Immunohistochemistry staining to evaluate the expression of Cdc45 in hepatocellular carcinoma (HCC) and adjacent tissues. (A, B) The expression of Cdc45 was absent or weak in the non-HCC tissue (2.565±1.410). (C, D) The expression of Cdc45 was obviously high in the HCC tissue (4.635±2.051). Cdc45 was detected in the nucleus and cytoplasm. Magnification, ×20 (A, C) and ×400 (B, D).
Figure 3. The Cdc45 expression levels for the in-house tissue microarrays. Immunohistochemistry staining to evaluate the expression of Cdc45 in hepatocellular carcinoma (HCC) and adjacent tissues. (A, B) The expression of Cdc45 was absent or weak in the non-HCC tissue (2.565±1.410). (C, D) The expression of Cdc45 was obviously high in the HCC tissue (4.635±2.051). Cdc45 was detected in the nucleus and cytoplasm. Magnification, ×20 (A, C) and ×400 (B, D). Figure 4. The co-expressed genes based on the Chen Liver microarray. NUDT1 and E2F1 were the top 2 genes on the list of co-expressed genes of CDC45 (r=0.713 and 0.675, respectively). The correlation index of other genes was 0.652.
Figure 4. The co-expressed genes based on the Chen Liver microarray. NUDT1 and E2F1 were the top 2 genes on the list of co-expressed genes of CDC45 (r=0.713 and 0.675, respectively). The correlation index of other genes was 0.652. Figure 5. The co-expressed genes based on the Wurmbach Liver microarray. All the correlation indexes of co-expressed genes of CDC45 were 0.666, except CCNE2 (r=0.757).
Figure 5. The co-expressed genes based on the Wurmbach Liver microarray. All the correlation indexes of co-expressed genes of CDC45 were 0.666, except CCNE2 (r=0.757). Figure 6. The co-expressed genes based on the Roessler Liver microarray. MCM5, CENPM, GINS2, ASF1B, E2F8, RNASEH2A, and CDKN2C were the top genes co-expressed with CDC45 in Roessler Liver.
Figure 6. The co-expressed genes based on the Roessler Liver microarray. MCM5, CENPM, GINS2, ASF1B, E2F8, RNASEH2A, and CDKN2C were the top genes co-expressed with CDC45 in Roessler Liver. Figure 7. The clinical implications of CDC45 genetic alterations. (A) The oncoprint of CDC45 in 360 patients/complete samples from 442 cases of hepatocellular carcinoma (HCC). CDC45 was altered in 34 (9%) cases. Patients with the CDC45 genetic alteration were more likely to have poor lifespan (B, for overall survival; C, for disease-free survival).
Figure 7. The clinical implications of CDC45 genetic alterations. (A) The oncoprint of CDC45 in 360 patients/complete samples from 442 cases of hepatocellular carcinoma (HCC). CDC45 was altered in 34 (9%) cases. Patients with the CDC45 genetic alteration were more likely to have poor lifespan (B, for overall survival; C, for disease-free survival). Figure 8. The CDC45 genetic alterations in multiple HCC datasets. (A) The oncoprint of CDC45 in 1089 cases of hepatocellular carcinoma (HCC) from multiple datasets involved The Cancer Genome Atlas (TCGA), AMC, Inserm, MSK, and RIKEN. CDC45 was altered in 9 (1%) of the patients. (B) The alteration frequency of CDC45 in HCC and mutation was the most common alternate forms, especially in liver (AMC) and HCC (Inserm, 2015) studies.
Figure 8. The CDC45 genetic alterations in multiple HCC datasets. (A) The oncoprint of CDC45 in 1089 cases of hepatocellular carcinoma (HCC) from multiple datasets involved The Cancer Genome Atlas (TCGA), AMC, Inserm, MSK, and RIKEN. CDC45 was altered in 9 (1%) of the patients. (B) The alteration frequency of CDC45 in HCC and mutation was the most common alternate forms, especially in liver (AMC) and HCC (Inserm, 2015) studies. Figure 9. The prognostic values of CDC45 genetic alterations in hepatocellular carcinoma (HCC). The survival time of altered group was less than the unaltered group, both for overall survival (A) and disease-free survival (B).
Figure 9. The prognostic values of CDC45 genetic alterations in hepatocellular carcinoma (HCC). The survival time of altered group was less than the unaltered group, both for overall survival (A) and disease-free survival (B). In Press
05 Mar 2024 : Clinical Research
Effects of Thermal Insulation on Recovery and Comfort of Patients Undergoing Holmium Laser LithotripsyMed Sci Monit In Press; DOI: 10.12659/MSM.942836
05 Mar 2024 : Clinical Research
Role of Critical Shoulder Angle in Degenerative Type Rotator Cuff Tears: A Turkish Cohort StudyMed Sci Monit In Press; DOI: 10.12659/MSM.943703
06 Mar 2024 : Clinical Research
Comparison of Outcomes between Single-Level and Double-Level Corpectomy in Thoracolumbar Reconstruction: A ...Med Sci Monit In Press; DOI: 10.12659/MSM.943797
21 Mar 2024 : Meta-Analysis
Economic Evaluation of COVID-19 Screening Tests and Surveillance Strategies in Low-Income, Middle-Income, a...Med Sci Monit In Press; DOI: 10.12659/MSM.943863
Most Viewed Current Articles
17 Jan 2024 : Review article
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
14 Dec 2022 : Clinical Research
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387
01 Jan 2022 : Editorial
Editorial: Current Status of Oral Antiviral Drug Treatments for SARS-CoV-2 Infection in Non-Hospitalized Pa...DOI :10.12659/MSM.935952
Med Sci Monit 2022; 28:e935952








