13 January 2021: Database Analysis
Systems Pharmacology-Based Identification of Mechanisms of Action of for the Treatment of Hepatocellular Carcinoma
Lan-Lan Wang1BE, Chen Liao2C, Xiao-Qiang Li3C, Rong Dai2C, Qing-Wei Ren4B, Hai-Long Shi1C, Xiao-Ping Wang1F, Xue-Song Feng1E, Xu Chao14AG*DOI: 10.12659/MSM.927624
Med Sci Monit 2021; 27:e927624
Abstract
BACKGROUND: Traditional Chinese medicine has widely used Bolbostemma paniculatum to treat diseases, including cancer, but its underlying mechanisms remain unclear. The present study aimed to elucidate the potential pharmacological mechanisms of “Tu Bei Mu” (TBM), the Chinese name for Bolbostemmatis Rhizoma, the dry tuber of B. paniculatum, for the treatment of hepatocellular carcinoma (HCC).
MATERIAL AND METHODS: The active components and putative therapeutic targets of TBM were explored using SwissTargetPrediction, Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP), and Search Tool for Interactions of Chemicals (STITCH). The HCC-related target database was built using DrugBank, DisGeNet, Online Mendelian Inheritance in Man (OMIM), and Therapeutic Target Database (TTD). A protein–protein interaction network of the common targets was constructed, based on the matches between TBM potential targets and HCC-related targets, using Cytoscape software. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses of the cluster networks were used to elucidate the biological functions of TBM.
RESULTS: Pharmacological network diagrams of the TBM compound-target network and HCC-related target network were successfully constructed. A total of 22 active components, 191 predicted biological targets of TBM, and 3775 HCC-related targets were identified. Through construction of an HCC-related target database and a protein–protein interaction network of the common targets, TBM was predicted to be effective in treating HCC mainly through the PI3K-Akt, HIF-1, p53, and PPAR signaling pathways.
CONCLUSIONS: The PI3K/Akt, HIF1, p53, and PPAR pathways may play vital roles in TBM treatment of HCC. Also, the potential anti-cancer effect of TBM on HCC appears to stem from the synergetic effect of multiple targets and mechanisms.
Keywords: Carcinoma, Hepatocellular, Drugs, Chinese Herbal, Pharmacology, Databases, Chemical, Databases, Genetic, Liver Neoplasms, Protein Interaction Maps, systems biology
Background
Hepatocellular Carcinoma (HCC) is the sixth most common cancer and fourth most common cause of cancer deaths worldwide [1]. Chronic hepatitis C virus (HCV) infection, chronic hepatitis B virus (HBV) infection, nonalcoholic steatohepatitis, and alcohol abuse are the major risk factors for HCC [2]. Also, obesity, associated with type 2 diabetes, has been shown to increase the risk of HCC [3]. Unfortunately, despite advancements in technology and treatment, HCC remains among the few cancers with an increasing incidence. Its 5-year survival rate is only 18.1% [4]. Surgery is most often not suitable for HCC patients because the tumors are usually at a late stage when diagnosed [5]. Sorafenib has been approved by the Food and Drug Administration (FDA) for HCC treatment, but its survival benefits are reportedly limited and the response rate is also very low [6]. Therefore, more effective treatment strategies for HCC are urgently needed.
The clinical practice of traditional Chinese medicine (TCM) has played an important role in preventing and treating diseases in China for thousands of years. TCM treatments have multicomponent, multitarget, and multi-pathway characteristics [7]. Therefore, exploration of TCM and other natural products as effective and less toxic anti-cancer drugs presents great promise for preventing and treating liver cancer [8]. Bolbostemmatis Rhizoma (Chinese name “Tu Bei Mu”, TBM) is the dry tuber of
The network-regulatory effect of the cold- and heat-syndrome biomolecular network and cold and heat prescription system of TCM was firstly reported in 2007 [17]. Also, the term “network pharmacology” was proposed for the first time in the same year [18]. Based on the concept of systems biology, network pharmacology integrates many processes, such as high-throughput data integration, database retrieval, data mining, target prediction, and laboratory simulation [19]. In network pharmacology, the interaction between drugs and specific nodes in each network module is analyzed from a systematic point of view, including the “multicomponent-multitarget” mechanisms of TCM or compound prescription. TCM can also independently propose specific disease-related pathways known as “active component-target pathways”. Systematic and holistic characteristics of network pharmacology are consistent with the overall view of TCM and the principle of syndrome differentiation and treatment [20]. Therefore, network pharmacology serves to reveal the materialistic basis for the pharmacodynamic effects of TCM and their mechanisms.
In the present study, we used network pharmacology to explore the active components and putative therapeutic targets of TBM to elucidate potential pharmacological mechanisms utilized by TBM in the treatment of hepatocellular carcinoma (HCC).
Material and Methods
SCREENING OF CHEMICAL COMPOUNDS IN TBM:
The chemical components of TBM were obtained from the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP, http://www.tcmspw.com/tcmsp.php/), which is the largest pharmacological data platform for TCM [21]. Figure 1 shows the schematic of the network pharmacology of TBM for the treatment of HCC; 46 components of TBM were found.
SCREENING FOR THE ACTIVE CHEMICAL COMPONENTS OF TBM:
The oral administration route is best for TCM effects in the gastrointestinal tract. Oral bioactivity (OB) is used for determining the dosing regimen. OB signifies the percentage of orally administered drug that reaches the blood circulation [22]. Drug-likeness (DL) is a qualitative concept in drug design that describes and optimizes the pharmacokinetic and pharmaceutical properties of a prospective drug-like compound [23]. In this study, OB and DL were used to screen the bioactive chemical components of TBM. In the screening process, chemical components with OB ≥20% and DL ≥0.18 were chosen for further analysis [24]. Additionally, the putative targets of the active chemical components of TBM were identified using the TCMSP, SwissTargetPrediction, and STITCH. Chemical components lacking target information were excluded.
IDENTIFICATION OF TARGET GENES RELATED TO THE IDENTIFIED CHEMICAL COMPONENTS:
The relevant targets of the active chemical components of TBM were identified using the TCMSP, SwissTargetPrediction (http://www.swisstargetprediction.ch/), and STITCH (http://stitch.embl.de/) programs [25]. The TCMSP database offers comprehensive information about the chemical components present in herbs, including chemical structure, drug targets, and their relationships with diseases. Based on the probability value obtained from the cross-validation analysis in the SwissTargetPrediction database, the rank of the targets and the accuracy of the prediction were estimated. Furthermore, to investigate the mode of action of TBM, interactions among the identified protein targets were analyzed using the STITCH database. The target protein names of the TBM bioactive chemical components were acquired and the standard gene names and Uniprot ID of these target proteins were retrieved from the UniprotKB (http://www.uniprot.org/) database, using the species “Homo sapiens” [26].
COLLECTION OF TARGETS RELATED TO HCC:
Targets related to HCC were collected from the OMIM (http://omim.org/), Drugbank (http://www.drugbank.ca), and DisGeNET (https://www.disgenet.org/) databases. OMIM is an information platform for human genes and genetic diseases [27]. Drugbank offers information about the target genes of many FDA-approved drugs [28]. DisGeNET is among the largest publicly available discovery platforms containing collections of genes and their variants associated with human diseases [29]. The database was searched against the keyword “hepatocellular carcinoma” and a total of 3755 HCC-related targets were retrieved, excluding duplicates. The potential active chemical components of TBM were matched against the potential targets related to HCC. The overlapping targets were selected as the components of TBM that have potential bioactivity against HCC.
ANALYSIS OF NETWORK CONSTRUCTION AND CENTRAL NETWORK TOPOLOGY:
A protein–protein interaction (PPI) network was constructed by processing the common targets using the STRING (https://string-db.org/) program [30]. The PPI network was visualized using Cytoscape v3.7.2 software [31,32]. Central network evaluation is a topological method that defines the core central network [33]. Degree centrality (DC), betweenness centrality (BC), and closeness centrality were analyzed using the Network Analyzer plugin in Cytoscape. These were used to measure the topological importance of nodes in the network [34]. The threshold values of the hub nodes in the network analysis were the corresponding median values of each parameter.
CLUSTER ANALYSIS:
Cluster analysis is an important classification method and many cluster analysis algorithms have been reported for the PPI network in Cytoscape. In 1 study, the network stability of a module generated using the Molecular Complex Detection (MCODE) algorithm was found to be superior. Hence, MCODE was used to perform cluster analysis for the PPI network [35]. The criteria settings were as follows: node score cutoff=0.2; K-core=2; degree cutoff=2; and max depth=100.
ENRICHMENT ANALYSIS OF THE GO AND KEGG PATHWAYS:
Potential biological processes, cellular components, molecular functions, and pathways related to the overlapping TBM and HCC targets were explored. The Database for Annotation Visualization and Integrated Discovery (DAVID, http://david.nicifcrf.gov/), an online platform for high-throughput functional annotation bioinformatics, was used to carry out the functional annotation and enrichment analysis [36]. GO terms with Bonferroni-adjusted P-value <0.05 and KEGG pathways with P-value <0.05 were considered significant [37].
Results
TARGET NETWORK OF TBM COMPOUNDS:
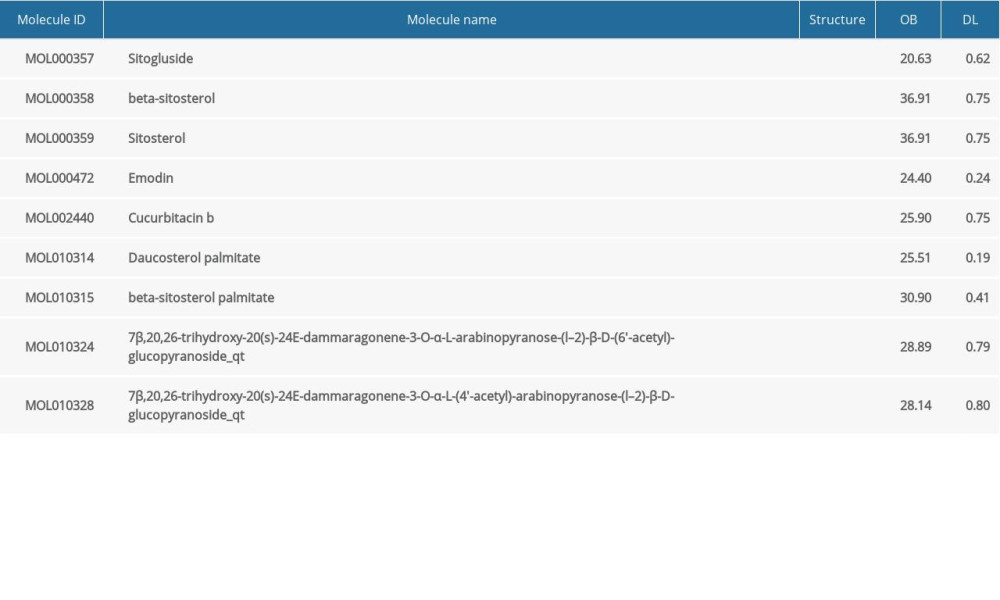
A total of 46 active TBM compounds were identified from the TCMSP database. These were further shortlisted based on the oral bioactivity (OB ≥20%) and drug-likeness (DL ≥0.18) standards. Out of the original 46, only 22 of the active TBM components fulfilled these criteria. Among these, 9 of the bioactive TBM components showed potential to bind to HCC targets while the other 13 did not correspond to any of the HCC targets. After rejecting the duplicates, a total of 191 putative targets for the 9 candidate compounds were identified. Their properties are shown in the Table 1.
TARGET GENES RELATED TO THE IDENTIFIED TBM CHEMICAL COMPONENTS:
The TBM compound-target network was constructed to elucidate the relationship between the compounds and the potential targets (Figure 2A). This network was composed of 200 nodes and 334 edges. Based on DC and BC, it was predicted that sitogluside, beta-sitosterol, sitosterol, and emodin are the principal active compounds that carry out the critical functions of TBM.
GO and KEGG pathway enrichment analyses were further performed to elucidate the characteristics of TBM-related targets. The GO analysis showed that most of the potential targets possess steroid-binding properties. Most of the target proteins were enriched in steroid-related processes, including the regulation of steroid hormone-mediated signaling pathways, drug response pathways, intracellular receptor signaling pathways, and in processes related to positive regulation of cell proliferation. These results suggest that TBM possesses multiple biological synergies (Figure 2B). The KEGG enrichment analysis revealed nearly 70 pathways (P<0.05) that could be predicted to be affected by TBM. The top 10 enriched pathways involved neuroactive ligand-receptor interaction, progesterone-mediated oocyte maturation, taste transduction, pathways associated with cancer, the sphingolipid signaling pathway, cell cycle regulation, prostate cancer, the PI3K-Akt signaling pathway, and the hepatitis C and FoxO signaling pathways (Figure 2C).
HCC-RELATED TARGET NETWORK:
In the present study, 3755 targets related to HCC in the genomic database of human diseases were selected for analysis. Among these, TBM and HCC had 104 common targets (Figure 3A, 3B). These common targets also had high confidence scores as per the PPIs generated using the STRING database (Figure 3C). The threshold values of the first screening were degree ≥30, closeness ≥0.554, and betweenness ≥141.982. Topological analysis indicated that TP53, CASP3, VEGFA, EGFR, EGF, TNF, STAT3, HSP90AA1, ESR1, PTGS2, AR, PPARG, ERBB2, MAPK14, NR3C1, and CXCR4 were the top shared targets.
GO analysis suggested that the majority of these 16 core targets have protein-binding functions and that they are enriched in the nucleus and cytoplasm (Figure 3D). KEGG enrichment analysis showed that these targets were enriched in processes related to the positive regulation of nitric oxide biosynthesis, gene expression and transcription from the RNA polymerase II promoter, and negative regulation of the apoptotic process (Figure 3E). These results indicate that various biological processes are involved in the multiple synergistic effects of TBM against HCC.
ENRICHMENT ANALYSIS OF THE 104 TARGETS:
The PPI network of TBM targets and HCC targets was analyzed using MCODE. The final central PPI network was then classified into 4 clusters (Figure 4A). The cluster modules that included targets with enriched biological functions and signaling pathways were used to analyze the integral regulation of TBM in HCC treatment (Figure 4B, 4C). The following GO terms were included: (1) peptidyl-tyrosine phosphorylation, nitric oxide biosynthetic process, and MAPK activity; (2) cell division, cell proliferation, and G2/M transition of the mitotic cell cycle; (3) cholesterol biosynthetic process, oxidation-reduction process, and isoprenoid biosynthetic process; and (4) cellular lipid metabolic process, steroid hormone-mediated signaling pathway, and transcription initiation from the RNA polymerase II promoter.
The KEGG enrichment analysis revealed that different signaling pathways were enriched in different modules (Figure 5). Module 1 was associated with pathways related to cancer, including the PI3K-Akt signaling pathway, hypoxia-inducible factor-1 (HIF-1) signaling pathway, and TNF signaling pathway. Module 2 was associated with the cell cycle and p53 signaling pathways involved in hepatitis B and hepatitis C. Module 3 was associated with steroid biosynthesis and metabolic pathways. Module 4 was associated with the PPAR signaling pathway and adipocytokine signaling pathway, including pathways associated with insulin resistance and nonalcoholic fatty liver disease.
KEGG data analysis also indicated that the key signaling pathways, such as the PI3K-Akt signaling pathway, the HIF-1 signaling pathway, the p53 signaling pathway, and the PPAR signaling pathway, might be the core pharmacological targets of TBM against HCC. Overall, the aforementioned analysis emphasizes a novel strategy for developing drugs against HCC.
Discussion
PI3K-AKT SIGNALING PATHWAY:
In several human malignant tumors, the PI3K/Akt/mTOR signaling pathway has been found to be aberrantly activated [43]. Inhibiting any node of the PI3K/Akt pathway can compensate for deficiencies in chemotherapy, radiotherapy, and hormone therapy. This synergistic effect has been closely related to apoptosis resulting from the selective inhibition of PI3K or Akt activity [44]. The PI3K-Akt signaling pathway has also been established in the occurrence and development of HCC. Although it does not play a role in apoptosis, it plays a critical role in cell growth, proliferation, adhesion ability, and expression of cellular adhesion molecules in HCC. Studies have shown that inhibitors of the PI3K signaling pathway increase sensitivity towards radiotherapy-induced tumor cell apoptosis [45]. Radiotherapy could activate the FAK-PI3K/Akt signaling pathway, which enhances the proliferation of radiotherapy-resistant, surviving HCC cells. TBM can inhibit the expression of caveolin-1(CAV-1), an oncogenic membrane protein, to initiate the PI3K/Akt signaling pathway, thus inhibiting the metastasis of cancer cells [46].
HIF-1 SIGNALING PATHWAY:
Hypoxia-inducible factor-1 (HIF-1) is a key transcription factor that is produced by tumor cells under hypoxic conditions to survive the hypoxic microenvironment. HIF-1 participates in energy metabolism, proliferation, apoptosis, invasion, metastasis, angiogenesis, and resistance to radiotherapy and chemotherapy by activating several downstream target genes in tumor cells [47]. Several studies have shown that changes in HIF-1 expression level are closely related to the proliferation and apoptosis of HCC cells. Additionally, it has been found that HIF-1 can promote cell proliferation through cyclin A and cyclin D [48]. HIF-1 is a direct transcriptional activator of the VEGF pathway under hypoxic conditions. It is reported that the anti-angiogenic effect of polypeptide extract from scorpion venom may be related to reduced levels of PI3K, HIF-1α, VEGF-A, and Akt [49]. Further, knockout of HIF-1α and IL-8 can inhibit angiogenesis in HCC [50]. HIF-1 can also affect the expression of EMT markers, such as cadherin E, cadherin N, and vimentin, by regulating the Snail factor to promote the invasion and metastasis of HCC cells [51]. Compared with other liver diseases, the expression level of HIF-1 is much higher in the serum of liver cancer patients. The HIF-1 serum level could be used as a new marker for the diagnosis and prognosis of liver cancer [52]. It has been reported that unregulated expression of HIF-1 provides a microenvironment conducive to tumors, and potentially increases the chance of recurrence of HCC after tumor resection [53]. Hence, HIF-1 could be used as an indicator of HCC prognosis, but such a use certainly requires further validation [54]. It has been found that reagents such as YC-1, a potential HIF-1 inhibitor, can act on targets by stimulating HIF-dependent P300 dissociation from HIF-1 [55]. The finding that siRNA-based silencing of HIF-1α significantly inhibited the proliferation of hepatoma in hypoxic CBRH-7919 mice through the PI3K/Akt signaling pathway led to the suggestion to focus on this pathway as a potential anti-cancer therapeutic strategy [56].
P53 SIGNALING PATHWAY:
Cell tumor antigen p53, an important tumor suppressor gene, encodes for the p53 protein, which is involved in the regulation of the cell cycle, DNA repair, signal transduction, and apoptosis [57,58]. TBM has been reported to initiate apoptosis through a variety of endogenous (mitochondrial) and exogenous (death receptor) pathways to produce broad-spectrum cytotoxic effects on a variety of tumor cells. TBM I has been suggested to induce the exogenous pathway of Fas/FasL/caspase-8 in hepatoma HepG2 cells [59]. Also, TBM has been shown to activate endogenous apoptosis by regulating the Bcl-2 protein [60]. TBM II has been reported to induce G2/M arrest in HepG2 cells. The growth of rabbit VX2 hepatoma was also significantly inhibited by embolization with TBM microcapsules (5.1 mg/kg) [61]. This study suggests that TBM may also inhibit the angiogenesis of cancer cells by regulating necessary proteins related to this process.
PPAR SIGNALING PATHWAY:
The PPAR pathway plays an important role in liver lipid metabolism by widely regulating the expression of proteins related to free fatty acid transport and β-oxidation. It also participates in the maintenance of glucose homeostasis [62]. It is involved in the regulation of many diseases, such as diabetes, obesity, and atherosis. However, its role in regulating cell growth, proliferation, and tumorigenesis is still controversial. PPAR-α, an important transcriptional regulatory factor, inhibits inflammation, but the mechanism underlying its regulatory effect in tumor proliferation, occurrence, and development is dubious [63]. It has been shown that endogenous PPAR-α agonists induce hepatocyte hypertrophy in rodents [64]. In transgenic mice, HCV core protein increases the level of circulating nonesterified fatty acids (NEFAs), which activate PPAR-α-mediated hepatogenesis [65]. These findings suggest that the endogenous ligand-activated PPAR-α signaling pathway could be involved in hepatocyte proliferation and carcinogenesis.
OTHER RELEVANT SIGNALING PATHWAYS:
Etiological studies of liver cancer, excluding cancers with hepatitis virus infection, suggest that diabetes can increase the risk of liver cancer by 2- to 3-fold [66,67]. Inflammation is central to the immune response in infection and injury. Chronic inflammation, however, aggravates the growth and development of malignant cells, which is closely associated with an increased risk of cancer [68]. TBM exerts an anti-inflammatory immune response. The order of strength of anti-inflammatory and antitumor activities was as follows: TBM III >TBM II >TBM I [69]. This suggests that TBM also plays an important role in regulating inflammation and immune responses. For instance, the hepatitis virus protein HBX plays an important role in the development of HCC by promoting the progression of hepatitis to HCC. Additionally, the enrichment analysis suggests that HBX may also affect other organ tumors, including the occurrence and development of liver cancer.
In general, the network pharmacological analysis revealed possible mechanisms of action of
Conclusions
The present study predicted the targets of TBM and explored mechanisms that could underlie potential anti-HCC effects. The PI3K/Akt, HIF-1, p53, and PPAR pathways may all play vital roles in TBM treatment of HCC. Our results suggest that the potential anti-cancer effect of TBM on HCC is based on the synergetic effect of multiple targets and mechanisms. However, these hypotheses still must be verified through further laboratory-based experiments.
Figures
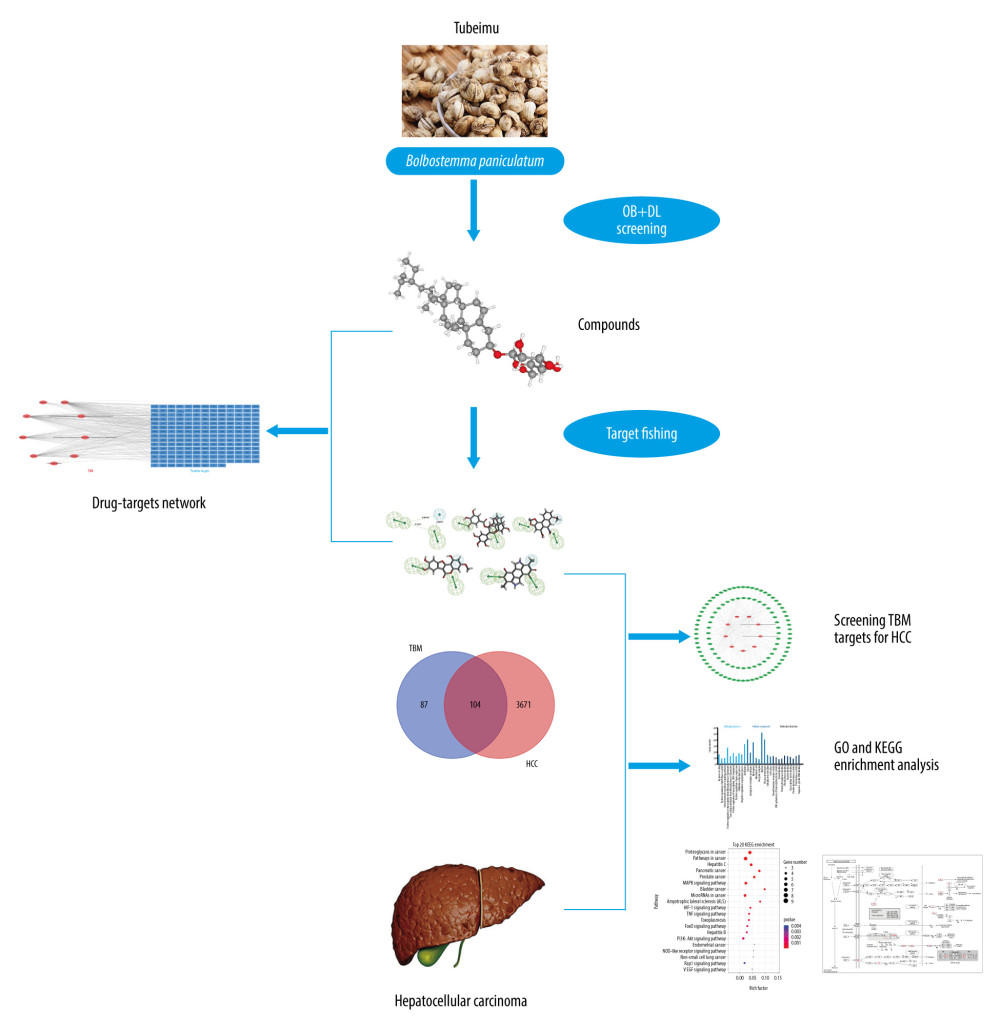 Figure 1. Schematic diagram of the network pharmacology-based analysis for investigating the potential mechanisms of TBM in treating HCC. TBM – Tu Bei Mu; HCC – hepatocellular carcinoma.
Figure 1. Schematic diagram of the network pharmacology-based analysis for investigating the potential mechanisms of TBM in treating HCC. TBM – Tu Bei Mu; HCC – hepatocellular carcinoma. 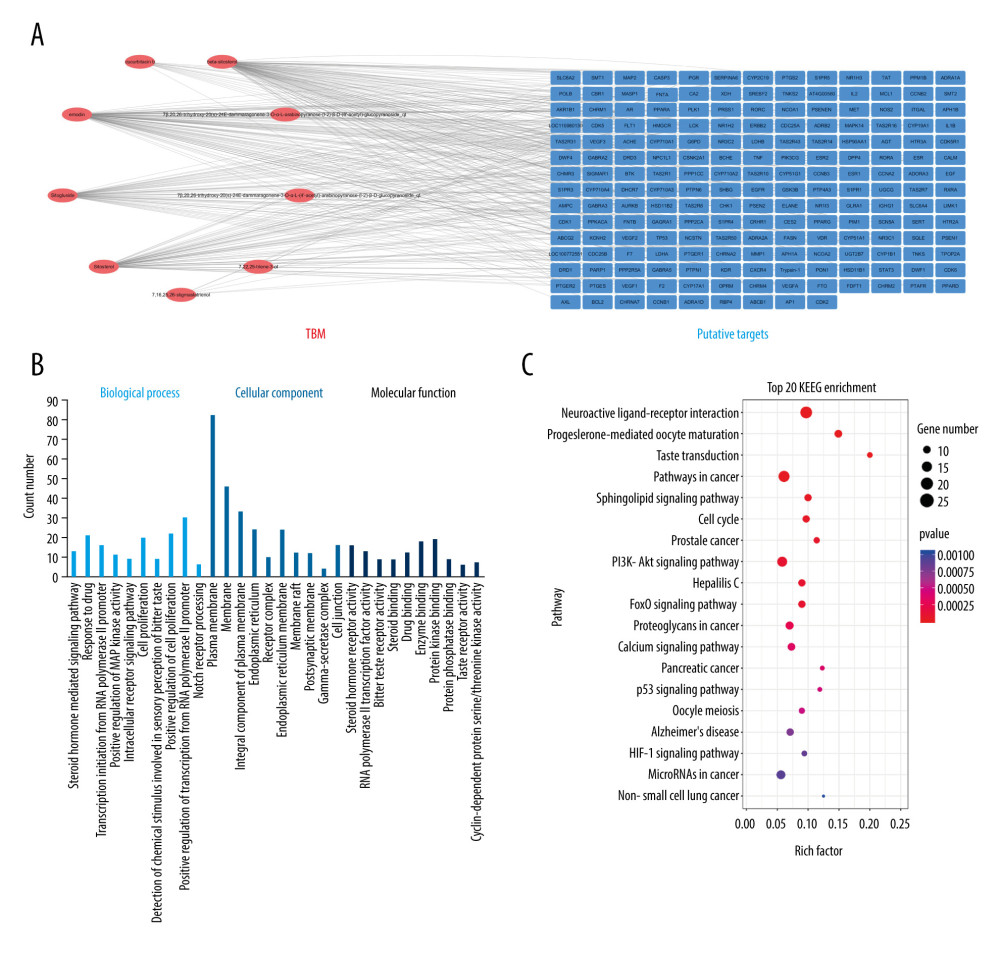 Figure 2. Characteristics of TBM-related targets clarified through GO and KEGG pathway enrichment analysis. (A) The component-target network of TBM. (B) GO enrichment analysis for potential targets of TBM. (C) KEGG enrichment analysis for potential targets of TBM. TBM – Tu Bei Mu; GO – Gene Ontology platform; KEGG – Kyoto Encyclopedia of Genes and Genomes platform.
Figure 2. Characteristics of TBM-related targets clarified through GO and KEGG pathway enrichment analysis. (A) The component-target network of TBM. (B) GO enrichment analysis for potential targets of TBM. (C) KEGG enrichment analysis for potential targets of TBM. TBM – Tu Bei Mu; GO – Gene Ontology platform; KEGG – Kyoto Encyclopedia of Genes and Genomes platform. 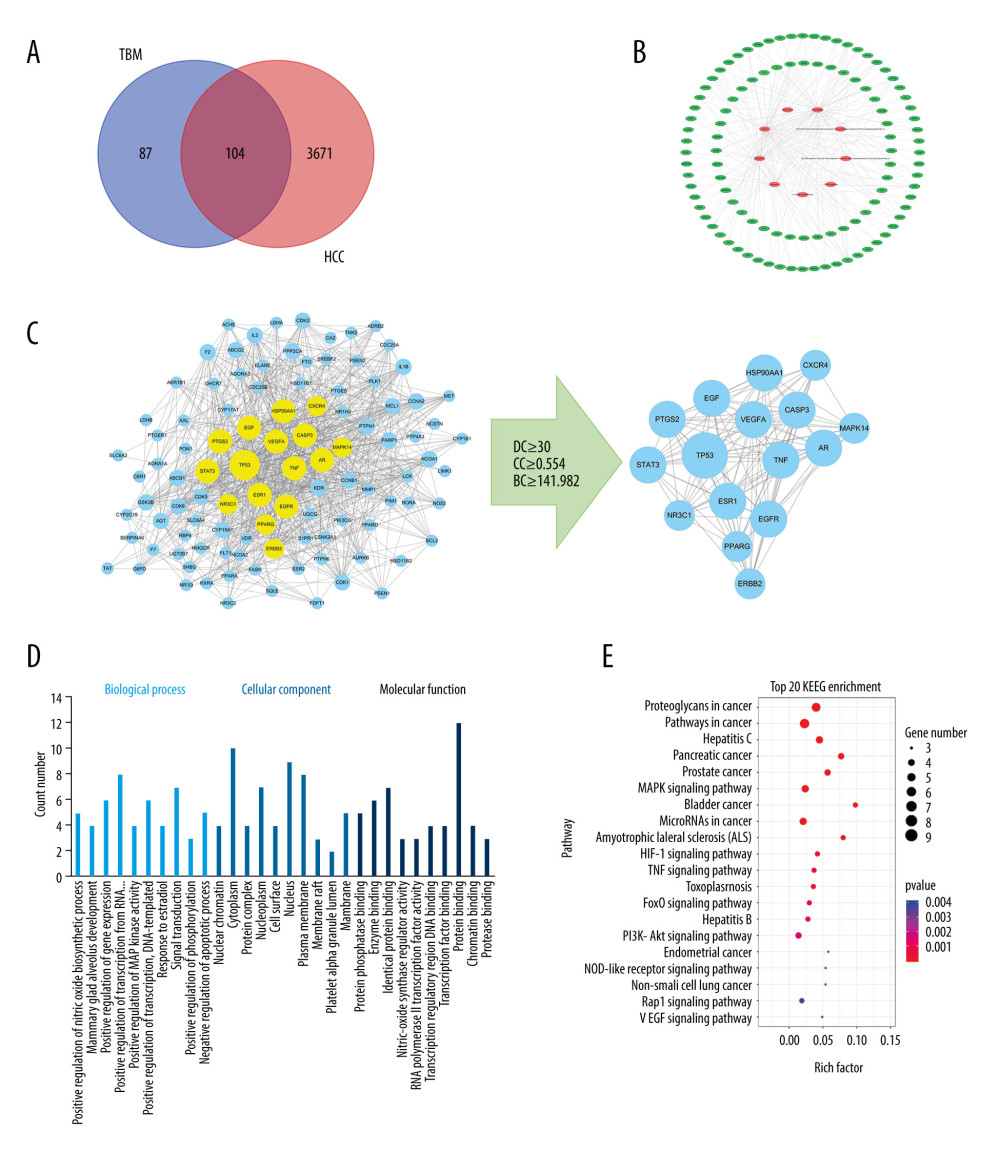 Figure 3. HCC-related target network. (A) 104 shared targets between TBM and HCC. (B) Component-target network of TBM for the treatment of HCC. The red circles represent components of TBM, and green hexagons represent common targets. (C) The process of topological screening of the PPI network. (D) GO enrichment analysis for 16 key targets. (E) KEGG enrichment analysis for 16 key targets. HCC – hepatocellular carcinoma; TBM – Tu Bei Mu; PPI – protein–protein interaction; GO – Gene Ontology platform; KEGG – Kyoto Encyclopedia of Genes and Genomes platform.
Figure 3. HCC-related target network. (A) 104 shared targets between TBM and HCC. (B) Component-target network of TBM for the treatment of HCC. The red circles represent components of TBM, and green hexagons represent common targets. (C) The process of topological screening of the PPI network. (D) GO enrichment analysis for 16 key targets. (E) KEGG enrichment analysis for 16 key targets. HCC – hepatocellular carcinoma; TBM – Tu Bei Mu; PPI – protein–protein interaction; GO – Gene Ontology platform; KEGG – Kyoto Encyclopedia of Genes and Genomes platform. 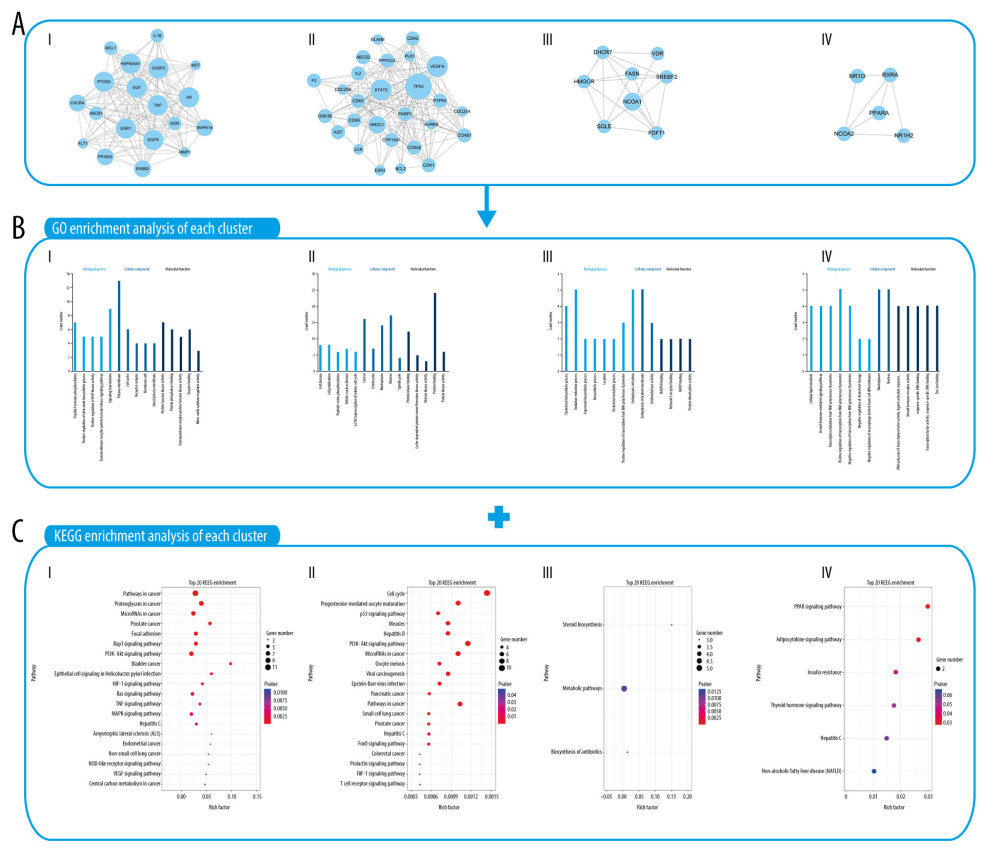 Figure 4. Enrichment analysis of the 104 TBM targets for the treatment of HCC. (A) Clusters of the final central PPI network. (B) GO enrichment analysis of each cluster. (C) KEGG pathway enrichment analysis of each cluster. TBM – Tu Bei Mu; HCC – hepatocellular carcinoma; PPI – protein–protein interaction; GO – Gene Ontology platform; KEGG – Kyoto Encyclopedia of Genes and Genomes platform.
Figure 4. Enrichment analysis of the 104 TBM targets for the treatment of HCC. (A) Clusters of the final central PPI network. (B) GO enrichment analysis of each cluster. (C) KEGG pathway enrichment analysis of each cluster. TBM – Tu Bei Mu; HCC – hepatocellular carcinoma; PPI – protein–protein interaction; GO – Gene Ontology platform; KEGG – Kyoto Encyclopedia of Genes and Genomes platform. 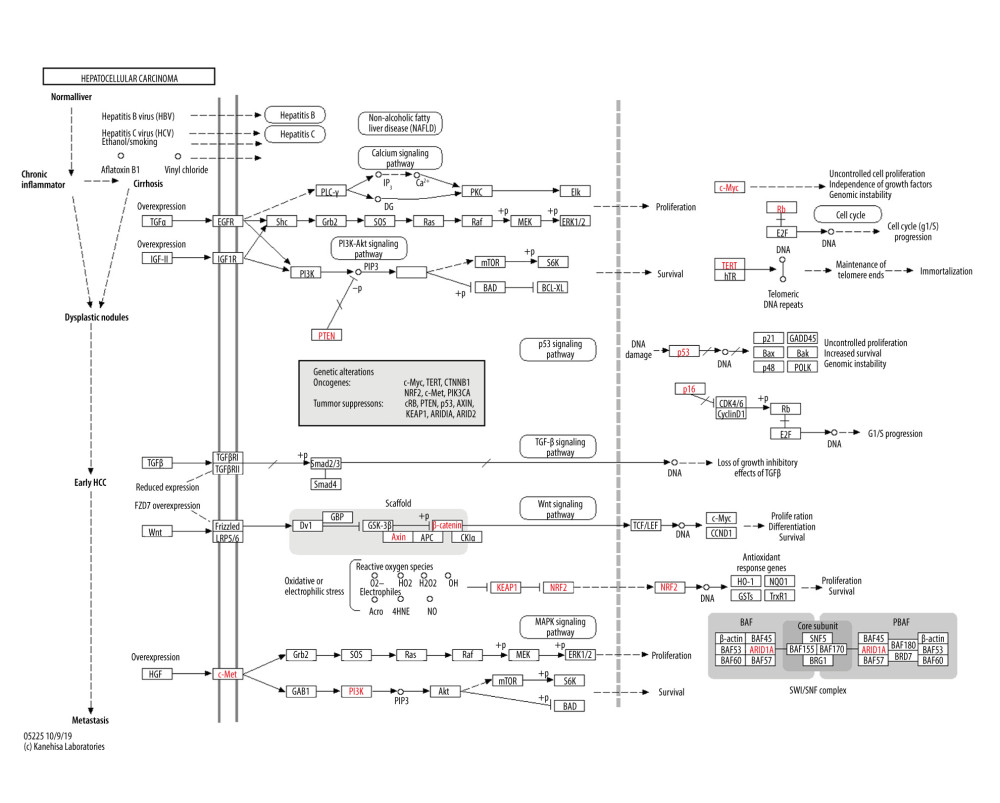 Figure 5. The predicted anticancer pathway of TBM. TBM – Tu Bei Mu.
Figure 5. The predicted anticancer pathway of TBM. TBM – Tu Bei Mu. References
1. Bray F, Ferlay J, Soerjomataram I, Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries: Cancer J Clin, 2018; 68(6); 394-424
2. Chimed T, Sandagdorj T, Znaor A, Cancer incidence and cancer control in Mongolia: Results from the National Cancer Registry 2008–12: Int J Cancer, 2017; 140(2); 302-9
3. Marengo A, Rosso C, Bugianesi E, Liver cancer: Connections with obesity, fatty liver, and cirrhosis: Annu Rev Med, 2016; 67; 103-17
4. Jemal A, Ward E-M, Johnson C-J, Annual Report to the Nation on the Status of Cancer, 1975–2014, featuring survival: J Natl Cancer Inst, 2017; 109 djx030
5. Chen C, Wang G, Mechanisms of hepatocellular carcinoma and challenges and opportunities for molecular targeted therapy: World J Hepatol, 2015; 7(15); 1964-70
6. Llovet JM, Focal gains of VEGFA: Candidate predictors of sorafenib response in hepatocellular carcinoma: Cancer Cell, 2014; 25(5); 560-2
7. Li S, Zhang B, Focal gains of VEGFA: candidate predictors of sorafenib response in hepatocellular carcinoma: Cancer Cell, 2014; 25(5); 560-62
8. Liao X, Bu Y, Jia Q, Traditional Chinese medicine as supportive care for the management of liver cancer: Past, present, and future: Genes Dis, 2019; 7(3); 370-79
9. Ail DA, Bhayekar P, Joshi A, Clinical and cytological spectrum of granulomatous mastitis and utility of FNAC in picking up tubercular mastitis: An eight-year study: J Clin Diagn Res, 2017; 11(3); EC45-49
10. Hu M, Zhao M, An C, Real-time imaging of apoptosis induction of human breast cancer cells by the traditional Chinese medicinal herb tubeimu: Anticancer Res, 2012; 32(7); 2509-14
11. Cheng G, Zhang Y, Zhang X, Tubeimoside V (1), a new cyclic bisdesmoside from tubers of Bolbostemma paniculatum, functions by inducing apoptosis in human glioblastoma U87MG cells: Bioorg Med Chem Lett, 2006; 16(17); 4575-80
12. Jiang SM, Xiao ZM, Song JG, Effects of BPF extracts on proliferation and mitochondria metabolism of human hepatoma SMMC-7721 cells: World Chin J Digestol, 2000; 8(3); 310-13
13. Yin Y, Chen W, Tang C: Food Chem Toxicol, 2011; 49(12); 3046-54
14. Zhao HB, Wang YS, Jiang XM: Chin Sci Bull, 2013; 58; 2572-79
15. Chao X, Zhao YY, Wei MH, Effects of tubeimoside-2 on proliferation and cell cycle of human hepatocellular carcinoma cell line HepG2: Jiangsu Med J, 2012; 38; 1740-42
16. Guo W, Liu Z, Dai S, Experimental study on the effect on the expression of VEGF and FGF2 after embolization by tubeimoside microcapsule: J Pract Radiol, 2008; 24; 1272-75
17. Li SFramework and practice of network-based studies for Chinese herbal formula: Zhong Xi Yi Jie He Xue Bao, 2007; 5(5); 489-93 [in Chinese]
18. Hopkins AL, Network pharmacology: Nat Biotechnol, 2007; 25(10); 1110-11
19. Zhang R, Zhu X, Bai H, Network pharmacology databases for traditional Chinese medicine: Review and assessment: Front Pharmacol, 2019; 10; 123
20. Berger SI, Iyengar R, Network analyses in systems pharmacology: Bioinformatics, 2009; 25(19); 2466-72
21. Ru J, Li P, Wang J, TCMSP: A database of systems pharmacology for drug discovery from herbal medicines: J Cheminform, 2014; 6; 13
22. Xu X, Zhang W, Huang C, A novel chemometric method for the prediction of human oral bioavailability: Int J Mol Sci, 2012; 13(6); 6964-82
23. Park K, Ko YJ, Durai P, Machine learning-based chemical binding similarity using evolutionary relationships of target genes: Nucleic Acids Res, 2019; 47(20); e128
24. Song W, Ni S, Fu Y, Uncovering the mechanism of Maxing Ganshi Decoction on asthma from a systematic perspective: A network pharmacology study: Sci Rep, 2018; 8(1); 17362
25. Kuhn M, von Mering C, Campillos M, STITCH: interaction networks of chemicals and proteins: Nucleic Acids Res, 2008; 36(Database issue); D684-88
26. Zaru R, Magrane M, Orchard S, Challenges in the annotation of pseudoenzymes in databases: The UniProtKB approach: FEBS J, 2020; 287(19); 4114-27
27. Amberger JS, Bocchini CA, Scott AF, OMIM.org: Leveraging knowledge across phenotype-gene relationships: Nucleic Acids Res, 2019; 47(D1); D1038-43
28. Probst D, Reymond JL, Exploring DrugBank in virtual reality chemical space: J Chem Inf Model, 2018; 58(9); 1731-35
29. Piñero J, Ramírez-Anguita JM, Saüch-Pitarch J, The DisGeNET knowledge platform for disease genomics: 2019 update: Nucleic Acids Res, 2020; 48(D1); D845-55
30. Crosara KTB, Moffa EB, Xiao Y: J Proteomics, 2018; 171; 87-94
31. Kohl M, Wiese S, Warscheid B, Cytoscape: Software for visualization and analysis of biological networks: Methods Mol Biol, 2011; 696; 291-303
32. Doncheva NT, Morris JH, Gorodkin J, Cytoscape StringApp: Network analysis and visualization of proteomics data: J Proteome Res, 2019; 18(2); 623-32
33. Tang Y, Li M, Wang J, CytoNCA: A cytoscape plugin for centrality analysis and evaluation of protein interaction networks: Biosystems, 2015; 127; 67-72
34. Li G, Li M, Wang J, United neighborhood closeness centrality and orthology for predicting essential proteins: IEEE/ACM Trans Comput Biol Bioinform, 2020; 17(4); 1451-58
35. Athanasios A, Charalampos V, Vasileios T, Protein–protein interaction (PPI) network: Recent advances in drug discovery: Curr Drug Metab, 2017; 18(1); 5-10
36. Huang DW, Sherman BT, Tan Q, DAVID bioinformatics resources: Expanded annotation database and novel algorithms to better extract biology from large gene lists: Nucleic Acids Res, 2007; 35(Web Server issue); W169-75
37. Chen L, Zhang YH, Wang SP, Prediction and analysis of essential genes using the enrichments of gene ontology and KEGG pathways: PLoS One, 2017; 12(9); e0184129
38. Uka K, Aikata H, Mori N, Combination therapy of oral fluoropyrimidine anti-cancer drug S-1 and interferon alpha for HCC patients with extrahepatic metastases: Oncology, 2008; 75(1–2); 8-16
39. Millimouno FM, Dong J, Yang L, Targeting apoptosis pathways in cancer and perspectives with natural compounds from mother nature: Cancer Prev Res (Phila), 2014; 7(11); 1081-107
40. Mehmood T, Maryam A, Ghramh HA, Deoxyelephantopin and isodeoxyelephantopin as potential anti-cancer agents with effects on multiple signaling pathways: Molecules, 2017; 22(6); 1013
41. Tang Y, Li W, Cao J, Bioassay-guided isolation and identification of cytotoxic compounds from Bolbostemma paniculatum: J Ethnopharmacol, 2015; 169; 18-23
42. Liu WY, Zhang WD, Chen HS, New triterpenoid saponins from bulbs of Bolbostemma paniculatum: Planta Med, 2004; 70(5); 458-64
43. Polivka J, Janku F, Molecular targets for cancer therapy in the PI3K/AKT/mTOR pathway: Pharmacol Ther, 2014; 142(2); 164-75
44. Khan KH, Yap TA, Yan L, Targeting the PI3K-AKT-mTOR signaling network in cancer: Chin J Cancer, 2013; 32(5); 253-65
45. Xiu P, Dong X, Dong X, Secretory clusterin contributes to oxaliplatin resistance by activating Akt pathway in hepatocellular carcinoma: Cancer Sci, 2013; 104(3); 375-82
46. Wang K, Zhu X, Chen Y, Tubeimoside V sensitizes human triple negative breast cancer MDA-MB-231 cells to anoikis via regulating caveolin-1-related signaling pathways: Arch Biochem Biophys, 2018; 646; 10-15
47. Yuan WC, Lee YR, Huang SF, A Cullin3-KLHL20 Ubiquitin ligase-dependent pathway targets PML to potentiate HIF-1 signaling and prostate cancer progression: Cancer Cell, 2011; 20(2); 214-28
48. Xu Z, Liu E, Peng C: Oncol Rep, 2012; 27(2); 573-78
49. Sui WW, Zhang WD, Wu LCStudy on the mechanism of polypeptide extract from scorpion venom on inhibition of angiogenesis of H22 hepatoma: Zhongguo Zhong Xi Yi Jie He Za Zhi, 2014; 34(5); 581-86 [in Chinese]
50. Choi SH, Kwon OJ, Park JY, Inhibition of tumour angiogenesis and growth by small hairpin HIF-1α and IL-8 in hepatocellular carcinoma: Liver Int, 2014; 34(4); 632-42
51. Zhang L, Huang G, Li X, Hypoxia induces epithelial-mesenchymal transition via activation of SNAI1 by hypoxia-inducible factor -1α in hepatocellular carcinoma: BMC Cancer, 2013; 13; 108
52. Li S, Yao D, Wang L, Expression characteristics of hypoxia-inducible factor-1α and its clinical values in diagnosis and prognosis of hepatocellular carcinoma: Hepat Mon, 2011; 11(10); 821-28
53. Simon F, Bockhorn M, Baba HA, Deregulation of HIF1-alpha and hypoxia-regulated pathways in hepatocellular carcinoma and corresponding non-malignant liver tissue – influence of a modulated host stroma on the prognosis of HCC: Langenbecks Arch Surg, 2010; 395(4); 395-405
54. Cao S, Yang S, Wu C, Protein expression of hypoxia-inducible factor-1 alpha and hepatocellular carcinoma: A systematic review with meta-analysis: Clin Res Hepatol Gastroenterol, 2014; 38(5); 598-603
55. Li SH, Shin DH, Chun YS, A novel mode of action of YC-1 in HIF inhibition: stimulation of FIH-dependent p300 dissociation from HIF-1α: Mol Cancer Ther, 2008; 7(12); 3729-38
56. Xu LF, Ni JY, Sun HL, Effects of hypoxia-inducible factor-1α silencing on the proliferation of CBRH-7919 hepatoma cells: World J Gastroenterol, 2013; 19(11); 1749-59
57. Joerger AC, Fersht AR, The p53 pathway: Origins, inactivation in cancer, and emerging therapeutic approaches: Annu Rev Biochem, 2016; 85; 375-404
58. Stegh AH, Targeting the p53 signaling pathway in cancer therapy – the promises, challenges and perils: Expert Opin Ther Targets, 2012; 16(1); 67-83
59. Yin Y, Chen W, Tang C: Food Chem Toxicol, 2011; 49(12); 3046-54
60. Wang Y, Deng L, Zhong H, Natural plant extract tubeimoside I promotes apoptosis-mediated cell death in cultured human hepatoma (HepG2) cells: Biol Pharm Bull, 2011; 34(6); 831-38
61. Rossi G, Salerno M, Granchi D, Change in FGF-2 circulating levels after arterial embolization in patients with bone metastases: Neoplasma, 2018; 65(2); 262-68
62. Rajamani K, Colman PG, Li LP, Effect of fenofibrate on amputation events in people with type 2 diabetes mellitus (FIELD study): A prespecified analysis of a randomised controlled trial: Lancet, 2009; 373(9677); 1780-88
63. Hou Y, Moreau F, Chadee K, PPARγ is an E3 ligase that induces the degradation of NFκB/p65: Nat Commun, 2012; 3; 1300
64. Kuwata K, Inoue K, Ichimura R, Constitutive active/androstane receptor, peroxisome proliferator-activated receptor α, and cytotoxicity are involved in oxadiazon-induced liver tumor development in mice: Food Chem Toxicol, 2016; 88; 75-86
65. Tanaka N, Moriya K, Kiyosawa K, Hepatitis C virus core protein induces spontaneous and persistent activation of peroxisome proliferator-activated receptor alpha in transgenic mice: Implications for HCV-associated hepatocarcinogenesis: Int J Cancer, 2008; 122(1); 124-31
66. Davila JA, Morgan RO, Shaib Y, Diabetes increases the risk of hepatocellular carcinoma in the United States: A population-based case control study: Gut, 2005; 54(4); 533-39
67. Wang C, Wang X, Gong G, Increased risk of hepatocellular carcinoma in patients with diabetes mellitus: A systematic review and meta-analysis of cohort studies: Int J Cancer, 2012; 130(7); 1639-48
68. Shacter E, Weitzman SA, Inflammation and cancer: Nature, 2002; 420(6917); 860-67
69. Yu TX, Ma RD, Yu LJ, Structure-activity relationship of tubeimosides in anti-inflammatory, antitumor, and antitumor-promoting effects: Acta Pharmacol Sin, 2001; 22(5); 463-68
Figures
 Figure 1. Schematic diagram of the network pharmacology-based analysis for investigating the potential mechanisms of TBM in treating HCC. TBM – Tu Bei Mu; HCC – hepatocellular carcinoma.
Figure 1. Schematic diagram of the network pharmacology-based analysis for investigating the potential mechanisms of TBM in treating HCC. TBM – Tu Bei Mu; HCC – hepatocellular carcinoma. Figure 2. Characteristics of TBM-related targets clarified through GO and KEGG pathway enrichment analysis. (A) The component-target network of TBM. (B) GO enrichment analysis for potential targets of TBM. (C) KEGG enrichment analysis for potential targets of TBM. TBM – Tu Bei Mu; GO – Gene Ontology platform; KEGG – Kyoto Encyclopedia of Genes and Genomes platform.
Figure 2. Characteristics of TBM-related targets clarified through GO and KEGG pathway enrichment analysis. (A) The component-target network of TBM. (B) GO enrichment analysis for potential targets of TBM. (C) KEGG enrichment analysis for potential targets of TBM. TBM – Tu Bei Mu; GO – Gene Ontology platform; KEGG – Kyoto Encyclopedia of Genes and Genomes platform. Figure 3. HCC-related target network. (A) 104 shared targets between TBM and HCC. (B) Component-target network of TBM for the treatment of HCC. The red circles represent components of TBM, and green hexagons represent common targets. (C) The process of topological screening of the PPI network. (D) GO enrichment analysis for 16 key targets. (E) KEGG enrichment analysis for 16 key targets. HCC – hepatocellular carcinoma; TBM – Tu Bei Mu; PPI – protein–protein interaction; GO – Gene Ontology platform; KEGG – Kyoto Encyclopedia of Genes and Genomes platform.
Figure 3. HCC-related target network. (A) 104 shared targets between TBM and HCC. (B) Component-target network of TBM for the treatment of HCC. The red circles represent components of TBM, and green hexagons represent common targets. (C) The process of topological screening of the PPI network. (D) GO enrichment analysis for 16 key targets. (E) KEGG enrichment analysis for 16 key targets. HCC – hepatocellular carcinoma; TBM – Tu Bei Mu; PPI – protein–protein interaction; GO – Gene Ontology platform; KEGG – Kyoto Encyclopedia of Genes and Genomes platform. Figure 4. Enrichment analysis of the 104 TBM targets for the treatment of HCC. (A) Clusters of the final central PPI network. (B) GO enrichment analysis of each cluster. (C) KEGG pathway enrichment analysis of each cluster. TBM – Tu Bei Mu; HCC – hepatocellular carcinoma; PPI – protein–protein interaction; GO – Gene Ontology platform; KEGG – Kyoto Encyclopedia of Genes and Genomes platform.
Figure 4. Enrichment analysis of the 104 TBM targets for the treatment of HCC. (A) Clusters of the final central PPI network. (B) GO enrichment analysis of each cluster. (C) KEGG pathway enrichment analysis of each cluster. TBM – Tu Bei Mu; HCC – hepatocellular carcinoma; PPI – protein–protein interaction; GO – Gene Ontology platform; KEGG – Kyoto Encyclopedia of Genes and Genomes platform. Figure 5. The predicted anticancer pathway of TBM. TBM – Tu Bei Mu.
Figure 5. The predicted anticancer pathway of TBM. TBM – Tu Bei Mu. In Press
08 Mar 2024 : Clinical Research
Evaluation of Foot Structure in Preschool Children Based on Body MassMed Sci Monit In Press; DOI: 10.12659/MSM.943765
15 Apr 2024 : Laboratory Research
The Role of Copper-Induced M2 Macrophage Polarization in Protecting Cartilage Matrix in OsteoarthritisMed Sci Monit In Press; DOI: 10.12659/MSM.943738
07 Mar 2024 : Clinical Research
Knowledge of and Attitudes Toward Clinical Trials: A Questionnaire-Based Study of 179 Male Third- and Fourt...Med Sci Monit In Press; DOI: 10.12659/MSM.943468
08 Mar 2024 : Animal Research
Modification of Experimental Model of Necrotizing Enterocolitis (NEC) in Rat Pups by Single Exposure to Hyp...Med Sci Monit In Press; DOI: 10.12659/MSM.943443
Most Viewed Current Articles
17 Jan 2024 : Review article
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
14 Dec 2022 : Clinical Research
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387
01 Jan 2022 : Editorial
Editorial: Current Status of Oral Antiviral Drug Treatments for SARS-CoV-2 Infection in Non-Hospitalized Pa...DOI :10.12659/MSM.935952
Med Sci Monit 2022; 28:e935952