24 July 2020: Database Analysis
Prognostic Value of Enolase Gene Family in Colon Cancer
Xiaohang Pan1AC, Huawen Wu1BF, Guofu Chen1EF, Wenhuan Li1AD*DOI: 10.12659/MSM.922980
Med Sci Monit 2020; 26:e922980
Abstract
BACKGROUND: Colorectal cancer (CRC), the most common gastrointestinal cancer, is associated with high mortality rates. Enolase is a major enzyme present in the glycolytic pathway. However, the functional significance of the enolase (ENO) gene family in the pathogenesis of CRC has been unclear.
MATERIAL AND METHODS: The data associated with 438 CRC patients from The Cancer Genome Atlas database were extracted for analysis. Survival analyses with Cox regression was performed to construct a prognostic signature. We investigated the processes that underlies the correlation between ENO genes and overall survival (OS) using gene set enrichment analysis (GSEA). We then developed a connectivity map to identify candidate target drugs for CRC.
RESULTS: The multivariate survival analysis showed that low expression of ENO2 and ENO3 had a significant correlation with longer OS. The joint-effects survival analysis indicated that the combined low expression of ENO2 and ENO3 was highly correlated with favorable OS. As indicated by the gene set enrichment analysis (GSEA), the ENO gene is involved in various biological pathways and has multiple roles. Potential pharmacological targets of ENO2 and ENO3 were constructed as well.
CONCLUSIONS: Low expression levels of both ENO2 and ENO3 were linked to a positive prognosis for CRC. Both ENO2 and ENO3 show promise as prognostic biomarkers for colon cancer patients.
Keywords: Colorectal Neoplasms, Phosphopyruvate Hydratase, Biomarkers, Tumor, Colonic Neoplasms, Databases, Genetic, Gene Expression Profiling, Proportional Hazards Models
Background
Colorectal cancer (CRC) is the most common gastrointestinal cancer and has high mortality. In the USA, the estimated incidence and mortality of CRC rank third among all cancers [1]. The 5-year relative survival rate for colorectal cancer patients is 65%. For patients with stage I or II disease, the 5-year relative survival rates are 91% and 82%, respectively, but the 5-year survival rate is only 12% for patients with stage IV disease. Moreover, the tumor stage has a strong association with CRC prognosis, and timely diagnosis and therapy improve overall survival (OS) rates [2].
Enolase is an important enzyme in the glycolytic pathway and is ubiquitous in organisms ranging from bacteria to mammals [3]. Enolase 1 (
Material and Methods
DATA SOURCE:
The medical data as well as the
ENO GENE FAMILY CORRELATION ANALYSIS AND BIOINFORMATICS ANALYSIS:
The co-expression analysis of ENO gene pathway and protein level expression was carried out with the use of GeneMANIA (www.genemania.org) [11], and functional bioinformatics analysis was performed in DAVID (david.ncifcrf.gov/tools.jsp) [12,13].
SURVIVAL ANALYSIS:
We assessed the prognosis of patients with CRC based on OS. The associations between the expression of ENO genes in CRC and the OS of patients were assessed using Kaplan-Meier analysis, log-rank test, and the Cox proportional hazards regression after adjusting for not age, sex, and TNM stage.
JOINT-EFFECTS SURVIVAL ANALYSIS:
Joint-effects analysis was used for the combination of significant ENO genes. The groups were developed by summarizing the chosen expression of ENO genes linked to better OS, worse OS, and other.
PROGNOSTIC RISK SCORE:
We formulated a prognostic risk score for the ENO2 and ENO3 genes in CRC. We used nomograms to predict 1-, 3-, 5-, and 10-year survival to evaluate the correlation between ENO genes and OS in CRC patients [14].
GENE SET ENRICHMENT ANALYSIS (GSEA):
The processes that underlie the correlation between the ENO genes and OS were investigated using GSEA. We performed biological pathway analysis in CRC with the ENO genes with the use of the reference c5 and c2 gene sets from the MSigDB, which made use of GSEA v.3.0 (http://software.broadinstitute.org/gsea/msigdb/index.jsp) [15]. The number of permutations was established at 1000. P<0.05 and FDR<0.25 were considered as having statistical significance.
PHARMACOLOGICAL TARGETS:
The genome-wide differentially expressed genes (DEGs), which include not just the upregulated but also the downregulated genes, together with the heatmaps and volcano plots, were attained with the use of edger [16]. Results with a fold change of >2 and P≤0.05 were used for analyses. Then, we chose target drugs from Connectivity Map (https://portals.broadinstitute.org/cmap/). In addition, the chemical compositions of related drugs were obtained from PubChem Compound (https://www.ncbi.nlm.nih.gov/pccompound/). The visualization of GO terms was done using DEGs with the use of BinGO. Thereafter, the enrichment analysis was carried on DEGs using DAVID.
STATISTICAL ANALYSIS:
SPSS v.25.0 software (IBM, Chicago, IL, USA) was used for statistical analyses. The calculation of OS was carried out using Kaplan-Meier analysis and log-rank test. The evaluation of the multivariate survival analysis was performed with log-rank P-values, hazard ratios (HR), and the calculation of the 95% percent confidence intervals (CIs) was done by Cox proportional hazards regression. P<0.05 was considered statistically significant.
Results
PATIENTS’ CLINICAL FEATURES:
The detailed clinical data attained from the TCGA concerning the 438 CRC patients are presented. Correlations between the clinical data and OS in the CRC patients are illustrated in Table 1 [17]. TNM stage had a significant association with OS (P<0.001; Table 1).
Scatter plots showing levels of ENO genes in CRC or normal colon tissue are shown in Figure 1. The median levels of ENO1, ENO2, and ENO3 were higher in CRC tissue than in normal colon tissue.
ENO GENE FAMILY CORRELATION ANALYSIS AND BIOINFORMATICS ANALYSIS:
Associations between expression of ENO genes in CRC were assed using Pearson correlation coefficients (Figure 2A). Figure 2B shows the pathway and co-expression prediction for ENO1, ENO2, ENO3, and ENO4. ENO gene family co-expression was assessed at the protein level (Figure 2C). The biological roles of the ENO genes were assessed based on the biological process, together with the molecular function and cellular component in GO pathway analysis. Findings associated with the KEGG pathway analysis are demonstrated in Figure 2D.
EFFECT OF DIFFERENTIAL ENO GENE EXPRESSION ON SURVIVAL:
Figure 3 shows the major results of univariate survival analysis. Low expression levels of ENO2 and ENO3 were significantly correlated with enhanced OS in CRC patients (P=0.003 and P<0.001, correspondingly). TNM stage was correlated with the prognosis of CRC patients (Table 1). Furthermore, the low expression levels of ENO2 (P=0.02) and ENO3(P<0.001) were associated with a longer OS (Table 2).
A joint-effects framework was constructed for the various cohorts based on the expression of ENO2 and ENO3 (Table 3). Low expression levels of ENO2 and ENO3 were significantly correlated with longer OS (P<0.001; Figure 4).
NOMOGRAM OF CRC PROGNOSTIC RISK SCORE MODEL:
The nomogram confirmed not that tumor stage and ENO2 and ENO3 expression in CRC predicted prognosis and contributed the majority of risk (range, 0–100 points) for poor OS. All of the variables were awarded points on the basis of Cox regression coefficients. The points were totaled, and the estimation of probability of survival made by drawing a vertical line (Figure 5).
GENE SET ENRICHMENT ANALYSIS (GSEA):
We performed the GSEA analysis to investigate the biological mechanisms underlying the effects of ENO2 and ENO3 overexpression. KEGG pathway analysis showed that overexpression of ENO2 was positively correlated with cell adhesion (Figure 6A), focal adhesion (Figure 6B), natural killer cells (Figure 6C), MAPK signaling pathway (Figure 6D), VEGF signaling pathway (Figure 6E), and cancer pathways (Figure 6F). GO enrichment analysis showed that overexpression of ENO2 had a positive correlation with cell adhesion (Figure 7A), as well as endothelial cell migration (Figure 7B), lymphocyte apoptotic process (Figure 7C), BMP signaling pathway (Figure 7D), ERK1 and ERK2 cascade (Figure 7E), and insulin-like growth factor receptor signaling pathway (Figure 7F).
PHARMACOLOGICAL TARGETS AND DRUGS:
We obtained DEGs with the use of edgeR. Pharmacological targets and drugs were attained from the Connectivity Map using the DEGs. The negatively correlated drugs constitute the latent pharmacological targets for ENO2 and ENO3 (Tables 4, 5). The heatmaps and volcano plots of these DEGs are demonstrated in Supplementary Figures 1–4. Supplementary Figures 5 and 6 show the chemical composition and the 2D structure of these latent target drugs. We performed enrichment analysis of the DEGs in DAVID. Supplementary Figure 7 and 8 show GO terms visualized by BinGO.
Discussion
We used data from TCGA to investigate correlations between the ENO gene expression levels in CRC, together with developing a risk score, including the medical factors as well as the expression patterns of ENO genes for the prediction of prognosis in patients with CRC. We found that expression levels of
Enolase was discovered in 1934 by Lohman and Mayerhof in the course of investigating the conversion of 3-phosphoglycerate to pyruvate in muscle extracts [18]. Enolase reaction has a major status in the metabolic pathway of fermentation generally, besides the glycolytic pathway, together with catalyzing the development of phosphoenolpyruvate from 2-phosphoglycerate, the second of the 2 high-power intermediates, generating the ATP in glycolysis [19. As indicated by the bioinformatics analysis of the current research work, the most evident molecular roles of
Enolase 2 (
Similarity to other malignant neoplasms, CRC is characterized by changes in the cell signaling and metabolic pathways, including energy metabolism [26].
In contrast to
Our study has certain limitations. First, the clinical information in the public databases was not detailed. Second, the patient data were from a single source. It is imperative to validate the prognostic significance of ENO genes in CRC by independent data containing full medical information. This was a bioinformatics investigation, and the majority results were created from the public database and bioinformatics analysis, lacking confirmation by
In spite of these constraints, this study is, to the best of our knowledge, the first to report that the downregulation of
Conclusions
We found that low expression levels of
Figures
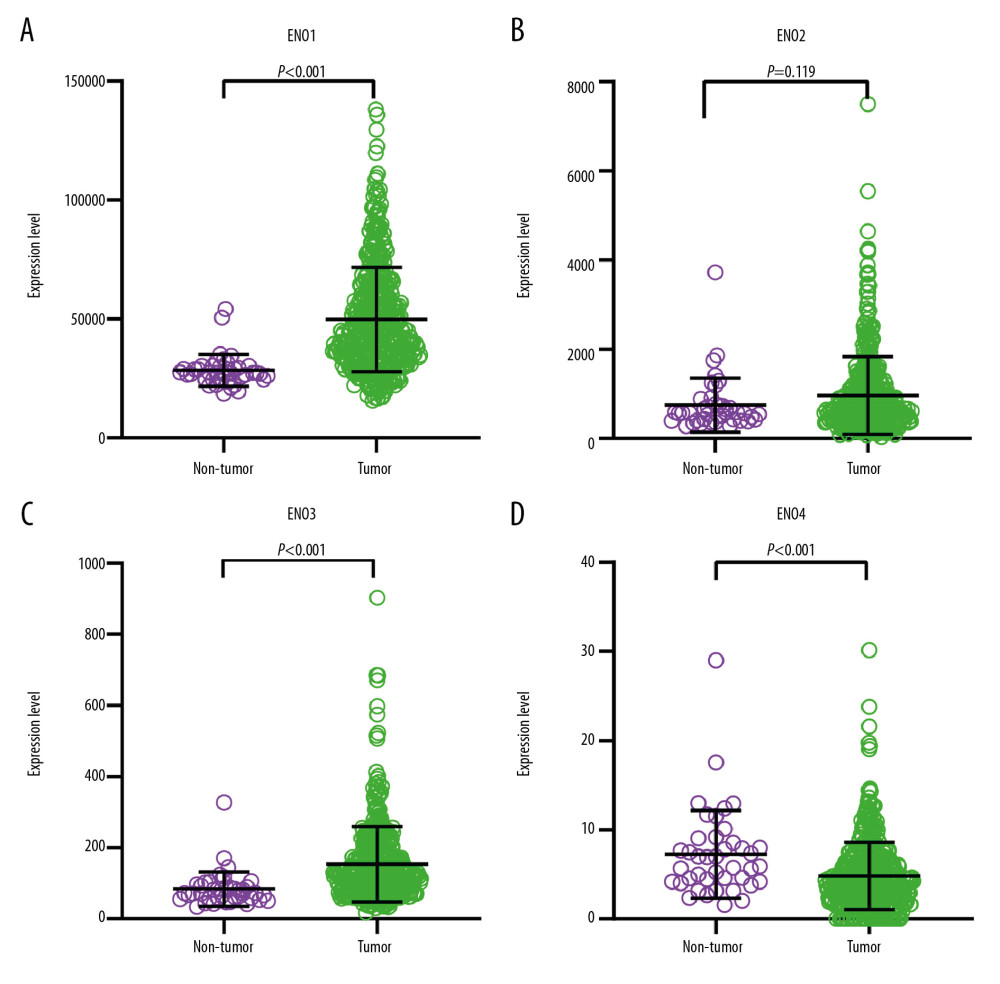 Figure 1. The scatter plots that show the expression profiles of ENO genes in CRC as well as normal colon tissue. (A) ENO1; (B) ENO2; (C) ENO3; (D) ENO4.
Figure 1. The scatter plots that show the expression profiles of ENO genes in CRC as well as normal colon tissue. (A) ENO1; (B) ENO2; (C) ENO3; (D) ENO4. 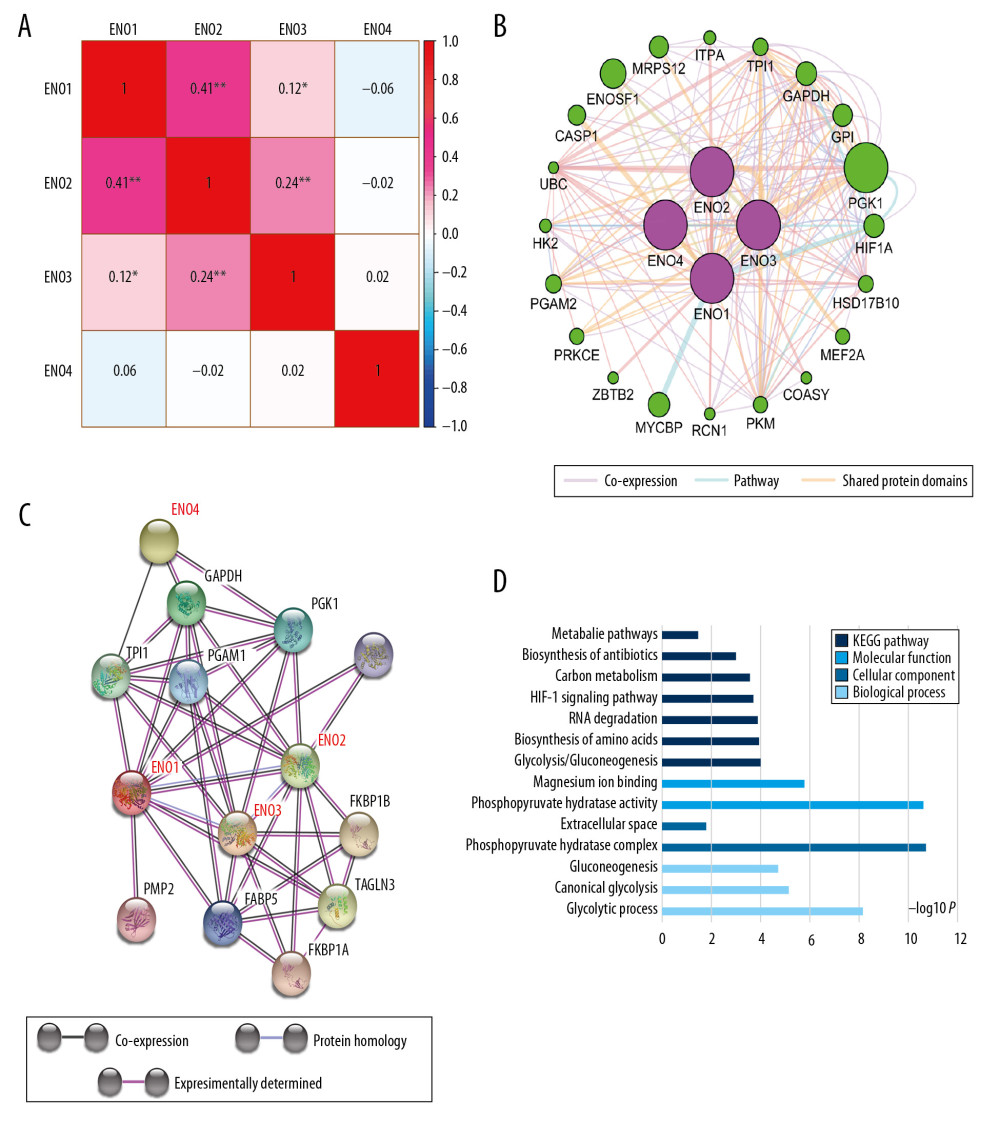 Figure 2. (A) Pearson’s correlation coefficients for ENO1, ENO2, ENO3 and ENO4 expression; (B) The gene interaction networks among ENO1, ENO2, ENO3 and ENO4; (C) The protein–protein interaction network of the ENO gene family; (D) GO pathway enrichment and KEGG pathway analysis carried out by DAVID.
Figure 2. (A) Pearson’s correlation coefficients for ENO1, ENO2, ENO3 and ENO4 expression; (B) The gene interaction networks among ENO1, ENO2, ENO3 and ENO4; (C) The protein–protein interaction network of the ENO gene family; (D) GO pathway enrichment and KEGG pathway analysis carried out by DAVID. 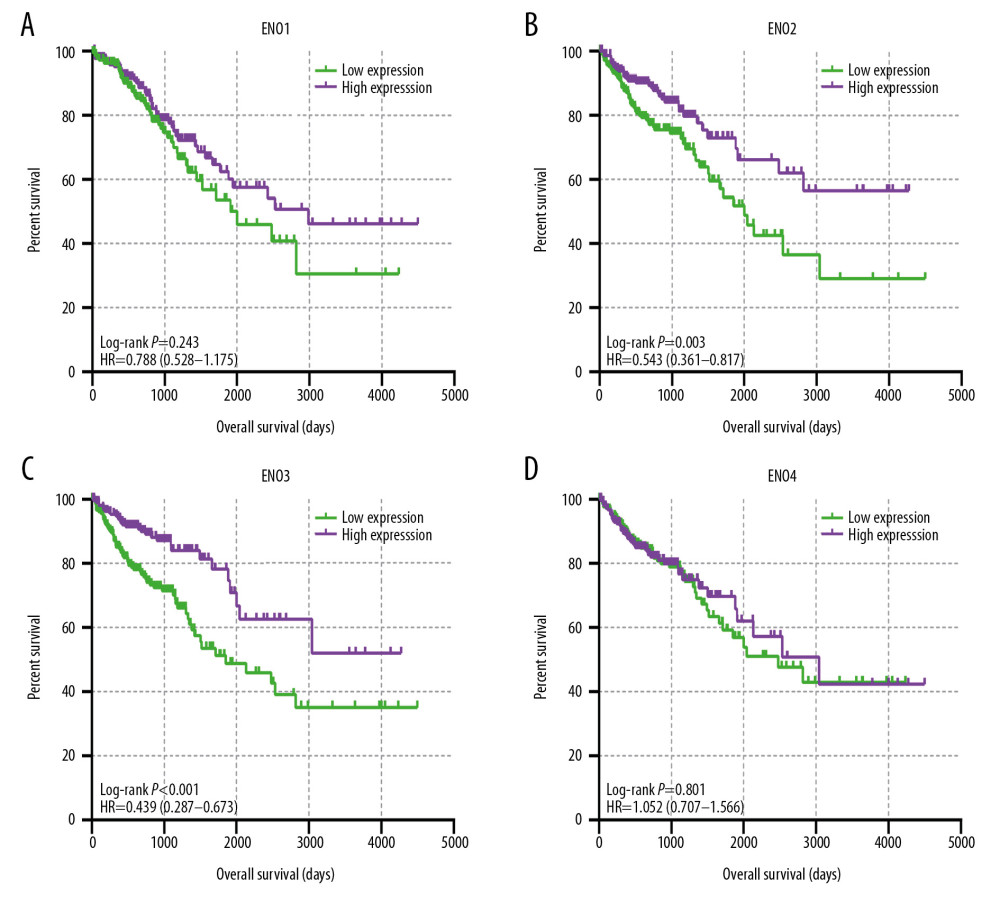 Figure 3. The prognostic significance of ENO expression for OS. (A–D) Kaplan-Meier survival curves concerning each of the colon cancer patients based on (A) ENO1, (B) ENO2, (C) ENO3, and (D) ENO4 expression (n=438).
Figure 3. The prognostic significance of ENO expression for OS. (A–D) Kaplan-Meier survival curves concerning each of the colon cancer patients based on (A) ENO1, (B) ENO2, (C) ENO3, and (D) ENO4 expression (n=438). 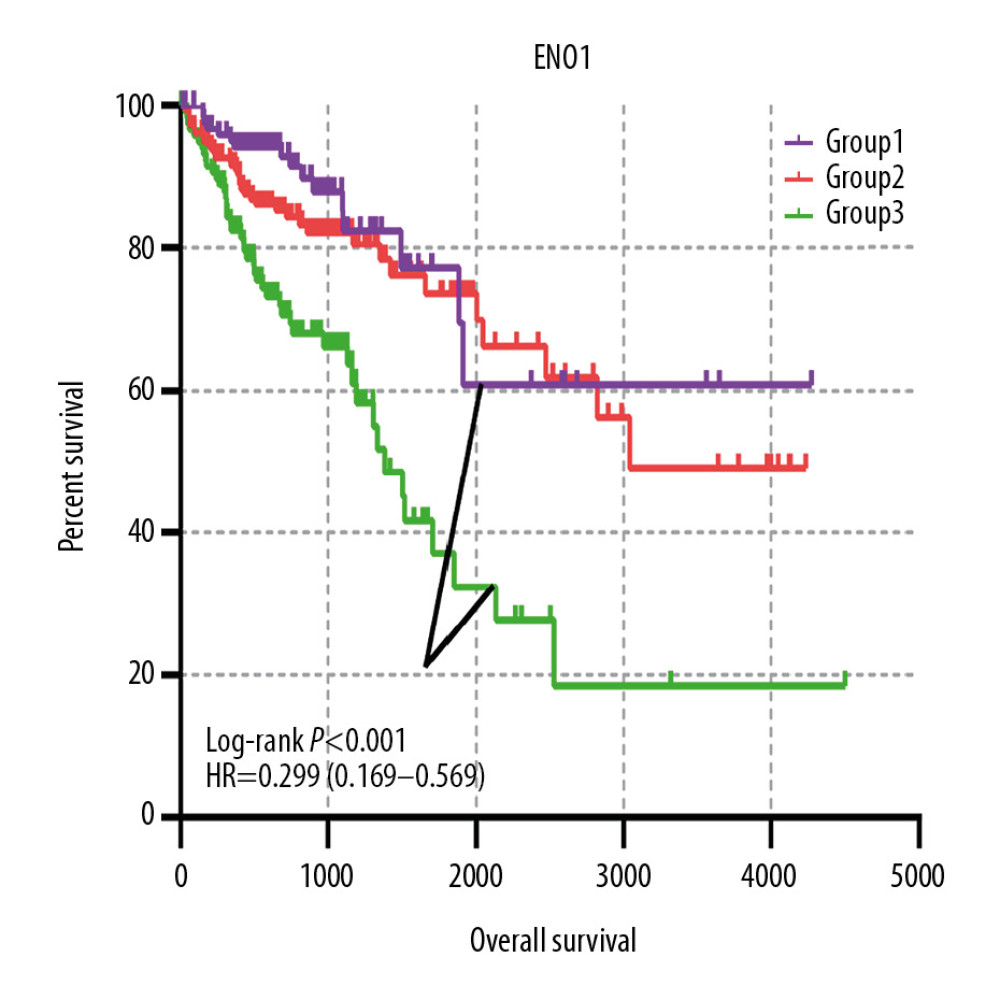 Figure 4. The joint-effects analysis of the influence of combined ENO gene expression on the OS with stratification on the basis of ENO2 and ENO3.
Figure 4. The joint-effects analysis of the influence of combined ENO gene expression on the OS with stratification on the basis of ENO2 and ENO3. 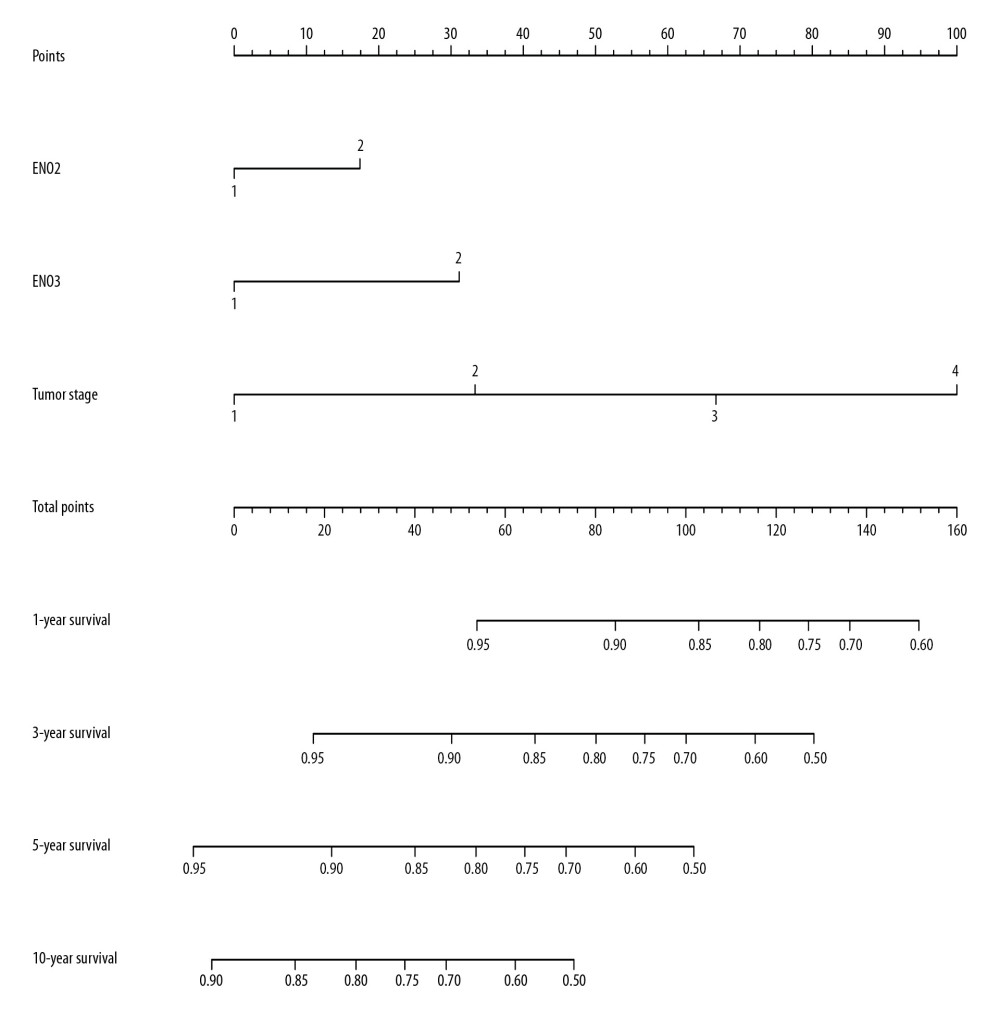 Figure 5. Nomogram for the prediction of OS in CRC with the use of ENO2, ENO3, and tumor stage.
Figure 5. Nomogram for the prediction of OS in CRC with the use of ENO2, ENO3, and tumor stage. 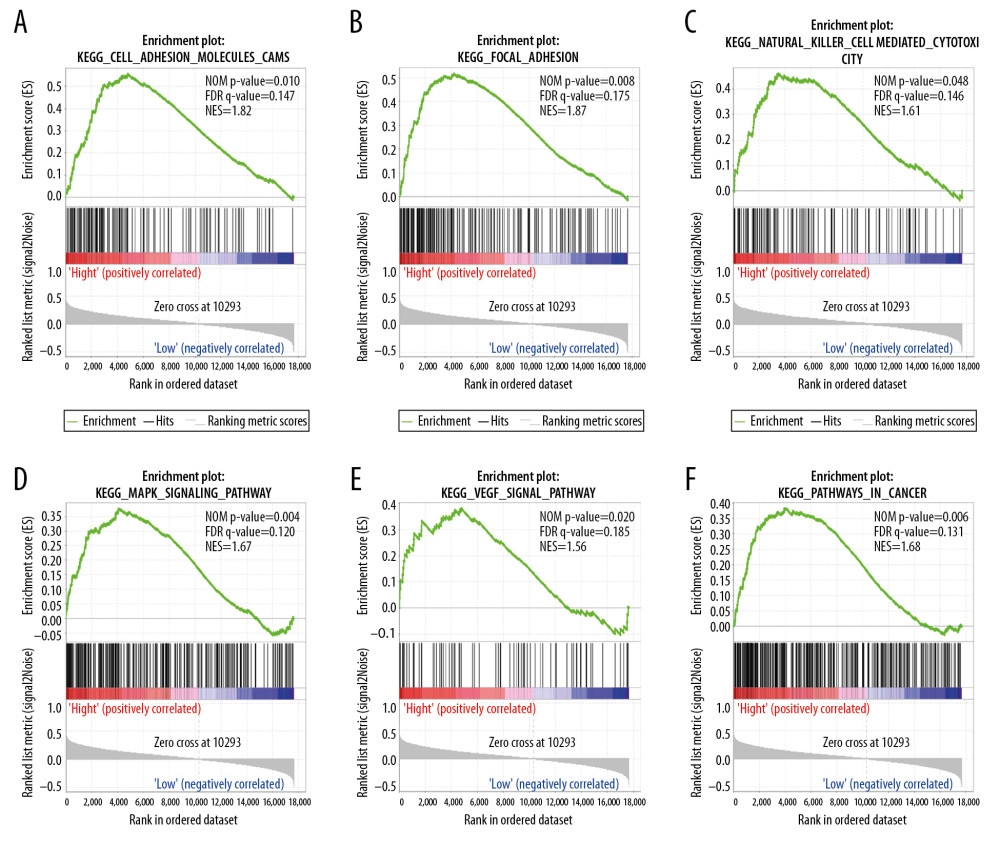 Figure 6. GSEA of ENO2 expressed in the colon cancer patients by the KEGG pathway analysis (A–F).
Figure 6. GSEA of ENO2 expressed in the colon cancer patients by the KEGG pathway analysis (A–F). 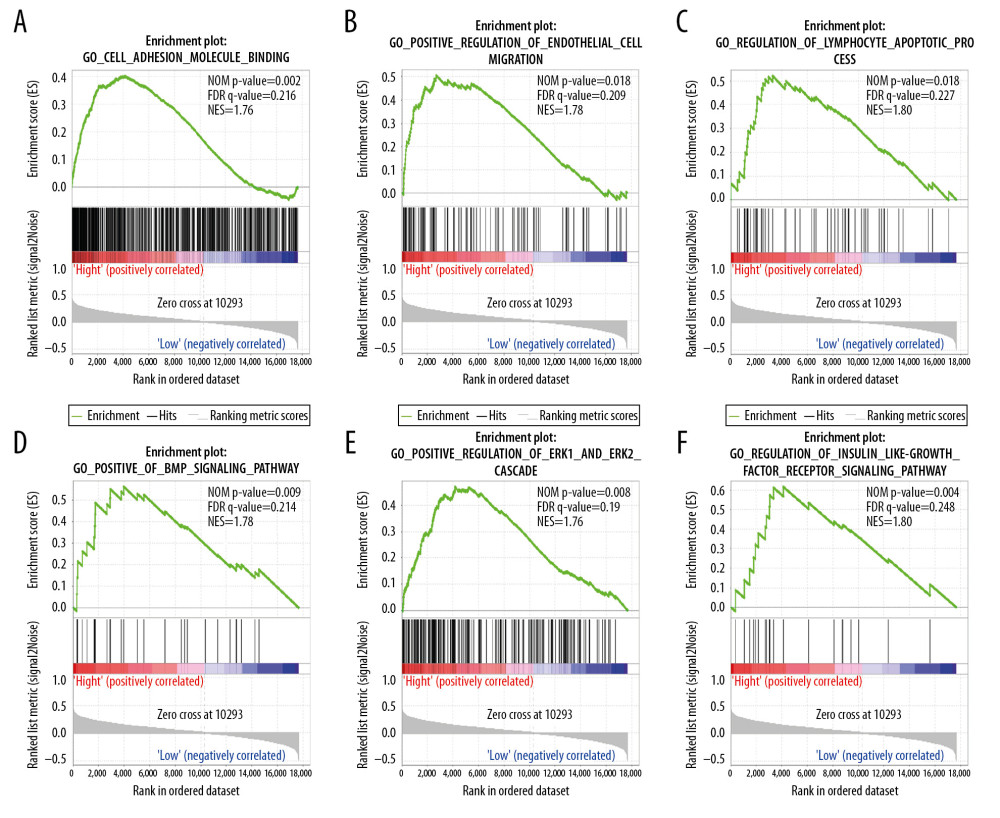 Figure 7. GSEA of ENO2 expressed in the colon cancer patients in accordance with the GO enrichment analysis (A–F).
Figure 7. GSEA of ENO2 expressed in the colon cancer patients in accordance with the GO enrichment analysis (A–F). References
1. Siegel RL, Miller KD, Jemal A, Cancer statistics, 2019: Cancer J Clin, 2019; 69; 7-34
2. Miller KD, Nogueira L, Mariotto AB, Cancer treatment and survivorship statistics, 2019: Cancer J Clin, 2019; 69; 363-85
3. Kang HJ, Jung SK, Kim SJ, Chung SJ, Structure of human alpha-enolase (hENO1), a multifunctional glycolytic enzyme: Acta Crystallogr D Biol Crystallogr, 2008; 64; 651-57
4. Ji M, Wang Z, Chen J, Up-regulated ENO1 promotes the bladder cancer cell growth and proliferation via regulating beta-catenin: Biosci Rep, 2019; 39 BSR20190503
5. Qiao H, Wang YF, Yuan WZ, Silencing of ENO1 by shRNA inhibits the proliferation of gastric cancer cells: Technol Cancer Res Treat, 2018; 17 1533033818784411
6. Zhan P, Wang Y, Zhao S, FBXW7 negatively regulates ENO1 expression and function in colorectal cancer: Lab Invest, 2015; 95; 995-1004
7. Xu L, Lina W, Xuejun Y, The diagnostic value of serum CEA, NSE and MMP-9 for on-small cell lung cancer: Open Med (Warsaw), 2016; 11; 59-62
8. Isgro MA, Bottoni P, Scatena R, Neuron-specific enolase as a biomarker: Biochemical and clinical aspects: Adv Exp Med Biol, 2015; 867; 125-43
9. Kong KW, Abdul Aziz A, Razali N: Peer J, 2016; 4; e2379
10. Higginbotham KS, Breyer JP, McReynolds KM: Cancer Epidemiol Biomarkers Prev, 2012; 21; 1565-73
11. Warde-Farley D, Donaldson SL, Comes O, The GeneMANIA prediction server: Biological network integration for gene prioritization and predicting gene function: Nucleic Acids Res, 2010; 38; W214-20
12. Huang da W, Sherman BT, Lempicki RA, Bioinformatics enrichment tools: Paths toward the comprehensive functional analysis of large gene lists: Nucleic Aacids Res, 2009; 37; 1-13
13. Huang da W, Sherman BT, Lempicki RA, Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources: Nat Protoc, 2009; 4; 44-57
14. Balachandran VP, Gonen M, Smith JJ, DeMatteo RP, Nomograms in oncology: More than meets the eye: Lancet Oncol, 2015; 16; e173-80
15. Subramanian A, Tamayo P, Mootha VK, Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles: Proc Natl Acad Sci USA, 2005; 102; 15545-50
16. Robinson MD, McCarthy DJ, Smyth GK, edgeR: A bioconductor package for differential expression analysis of digital gene expression data: Bioinformatics, 2010; 26; 139-40
17. Pan X, Wang Q, Xu C, Prognostic value of chloride channel accessory mRNA expression in colon cancer: Oncol Lett, 2019; 18; 2967-76
18. Pancholi V, Multifunctional alpha-enolase: Its role in diseases: Cell Mol Life Sci, 2001; 58; 902-20
19. Subramanian A, Miller DM, Structural analysis of alpha-enolase. Mapping the functional domains involved in down-regulation of the c-myc protooncogene: J Biol Chem, 2000; 275; 5958-65
20. Liu CC, Wang H, Wang WD, ENO2 promotes cell proliferation, glycolysis, and glucocorticoid-resistance in acute lymphoblastic leukemia: Cell Physiol Biochem, 2018; 46; 1525-35
21. Vizin T, Kos J, Gamma-enolase: A well-known tumour marker, with a less-known role in cancer: Radiol Oncol, 2015; 49; 217-26
22. Hafner A, Obermajer N, Kos J, γ-Enolase C-terminal peptide promotes cell survival and neurite outgrowth by activation of the PI3K/Akt and MAPK/ERK signalling pathways: Biochem J, 2012; 443; 439-50
23. Kaplan DR, Miller FD, Signal transduction by the neurotrophin receptors: Curr Opin Cell Biol, 1997; 9; 213-21
24. Jang S-M, Kim J-W, Kim C-H, p19ras Represses proliferation of non-small cell lung cancer possibly through interaction with Neuron-Specific Enolase (NSE): Cancer Lett, 2010; 289; 91-98
25. Soh MA, Garrett SH, Somji S, Arsenic, cadmium and neuron specific enolase (ENO2, gamma-enolase) expression in breast cancer: Cancer Cell Int, 2011; 11; 41
26. Snezhkina AV, Krasnov GS, Zaretsky AR, Differential expression of alternatively spliced transcripts related to energy metabolism in colorectal cancer: BMC Genomics, 2016; 17; 1011
27. Yeh CS, Wang JY, Chung FY, Significance of the glycolytic pathway and glycolysis related-genes in tumorigenesis of human colorectal cancers: Oncol Rep, 2008; 19; 81-91
28. Selga E, Morales C, Noe V, Role of caveolin 1, E-cadherin, Enolase 2 and PKCalpha on resistance to methotrexate in human HT29 colon cancer cells: BMC Med Genomics, 2008; 1; 35
29. Nasir SN, Abu N, Ab Mutalib NS, LOC285629 regulates cell proliferation and motility in colorectal cancer cells: Clin Transl Oncol, 2018; 20; 775-84
30. Zou M, Zhang PJ, Wen XY, A novel mixed integer programming for multi-biomarker panel identification by distinguishing malignant from benign colorectal tumors: Methods, 2015; 83; 3-17
31. Park C, Lee Y, Je S, Overexpression and selective anticancer efficacy of ENO3 in STK11 mutant lung cancers: Mol Cells, 2019; 42; 804-9
32. Liu ZK, Zhang RY, Yong YL, Identification of crucial genes based on expression profiles of hepatocellular carcinomas by bioinformatics analysis: Peer J, 2019; 7; e7436
Figures
 Figure 1. The scatter plots that show the expression profiles of ENO genes in CRC as well as normal colon tissue. (A) ENO1; (B) ENO2; (C) ENO3; (D) ENO4.
Figure 1. The scatter plots that show the expression profiles of ENO genes in CRC as well as normal colon tissue. (A) ENO1; (B) ENO2; (C) ENO3; (D) ENO4. Figure 2. (A) Pearson’s correlation coefficients for ENO1, ENO2, ENO3 and ENO4 expression; (B) The gene interaction networks among ENO1, ENO2, ENO3 and ENO4; (C) The protein–protein interaction network of the ENO gene family; (D) GO pathway enrichment and KEGG pathway analysis carried out by DAVID.
Figure 2. (A) Pearson’s correlation coefficients for ENO1, ENO2, ENO3 and ENO4 expression; (B) The gene interaction networks among ENO1, ENO2, ENO3 and ENO4; (C) The protein–protein interaction network of the ENO gene family; (D) GO pathway enrichment and KEGG pathway analysis carried out by DAVID. Figure 3. The prognostic significance of ENO expression for OS. (A–D) Kaplan-Meier survival curves concerning each of the colon cancer patients based on (A) ENO1, (B) ENO2, (C) ENO3, and (D) ENO4 expression (n=438).
Figure 3. The prognostic significance of ENO expression for OS. (A–D) Kaplan-Meier survival curves concerning each of the colon cancer patients based on (A) ENO1, (B) ENO2, (C) ENO3, and (D) ENO4 expression (n=438). Figure 4. The joint-effects analysis of the influence of combined ENO gene expression on the OS with stratification on the basis of ENO2 and ENO3.
Figure 4. The joint-effects analysis of the influence of combined ENO gene expression on the OS with stratification on the basis of ENO2 and ENO3. Figure 5. Nomogram for the prediction of OS in CRC with the use of ENO2, ENO3, and tumor stage.
Figure 5. Nomogram for the prediction of OS in CRC with the use of ENO2, ENO3, and tumor stage. Figure 6. GSEA of ENO2 expressed in the colon cancer patients by the KEGG pathway analysis (A–F).
Figure 6. GSEA of ENO2 expressed in the colon cancer patients by the KEGG pathway analysis (A–F). Figure 7. GSEA of ENO2 expressed in the colon cancer patients in accordance with the GO enrichment analysis (A–F).
Figure 7. GSEA of ENO2 expressed in the colon cancer patients in accordance with the GO enrichment analysis (A–F). Tables
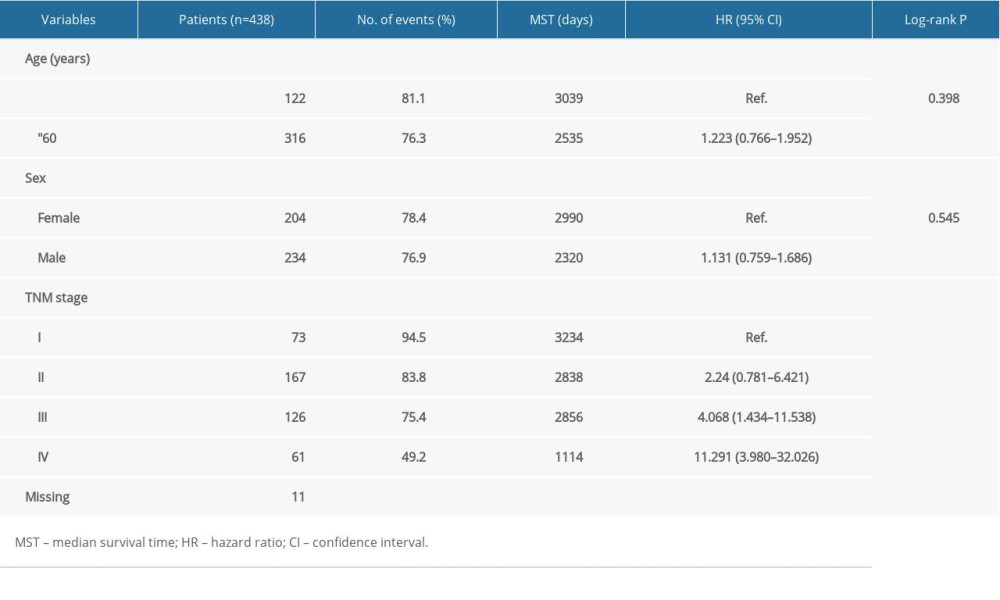 Table 1. Demographic and clinical data for 438 colon cancer patients.
Table 1. Demographic and clinical data for 438 colon cancer patients.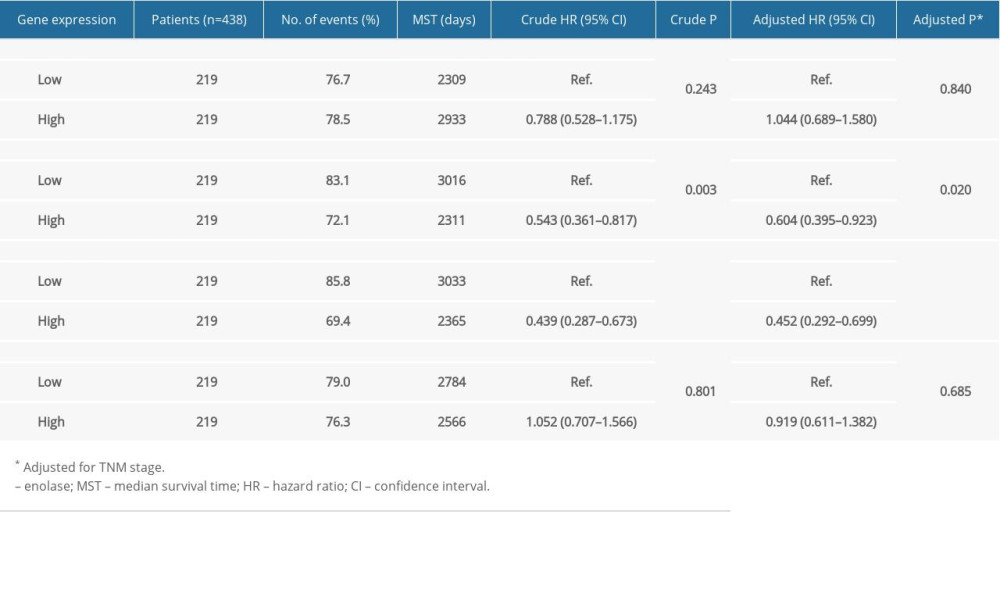 Table 2. Prognostic survival analysis of ENO family genes.
Table 2. Prognostic survival analysis of ENO family genes.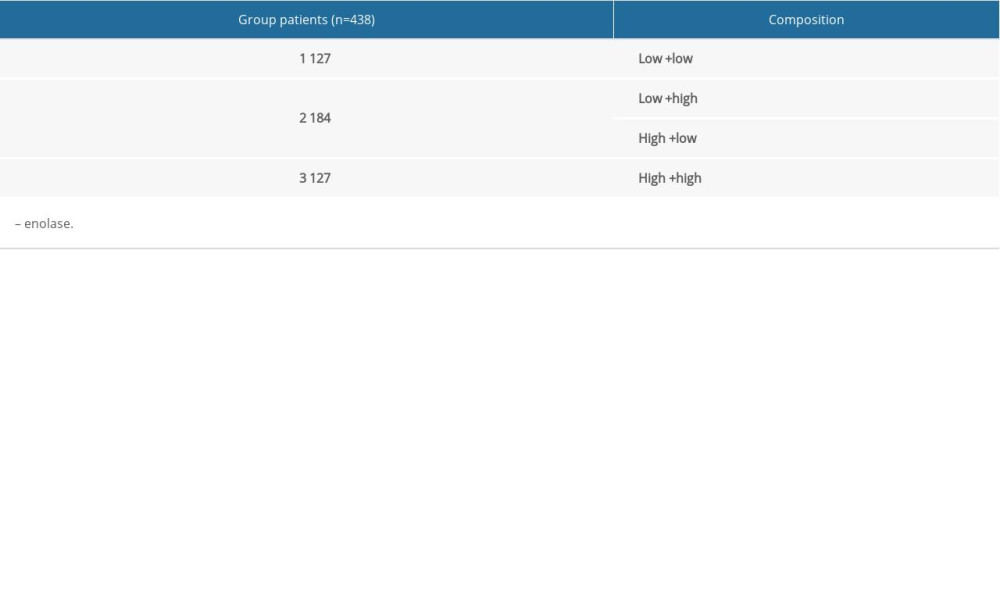 Table 3. Grouping according to ENO genes.
Table 3. Grouping according to ENO genes.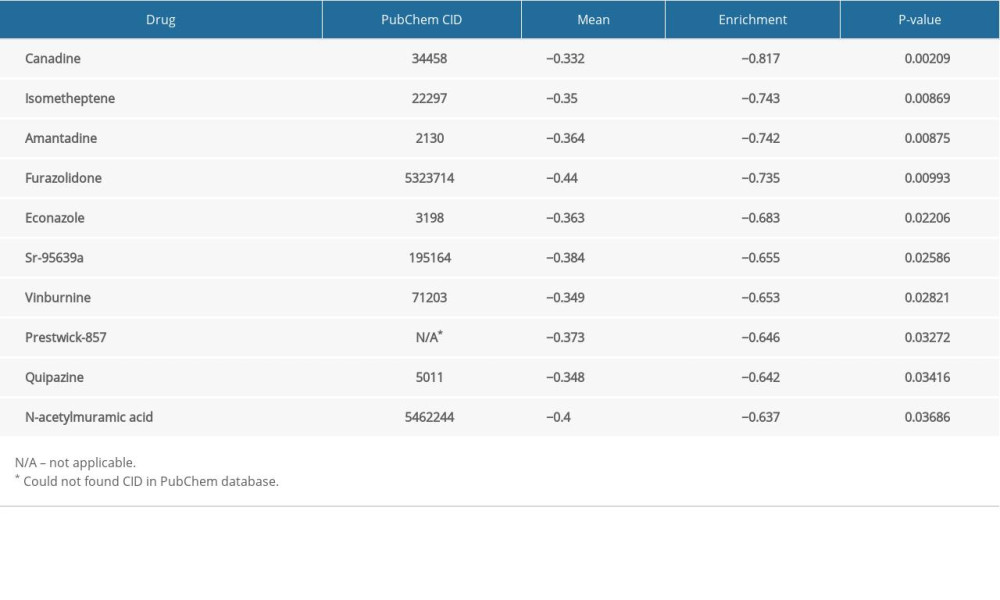 Table 4. Pharmacological targets and drug for ENO2.
Table 4. Pharmacological targets and drug for ENO2.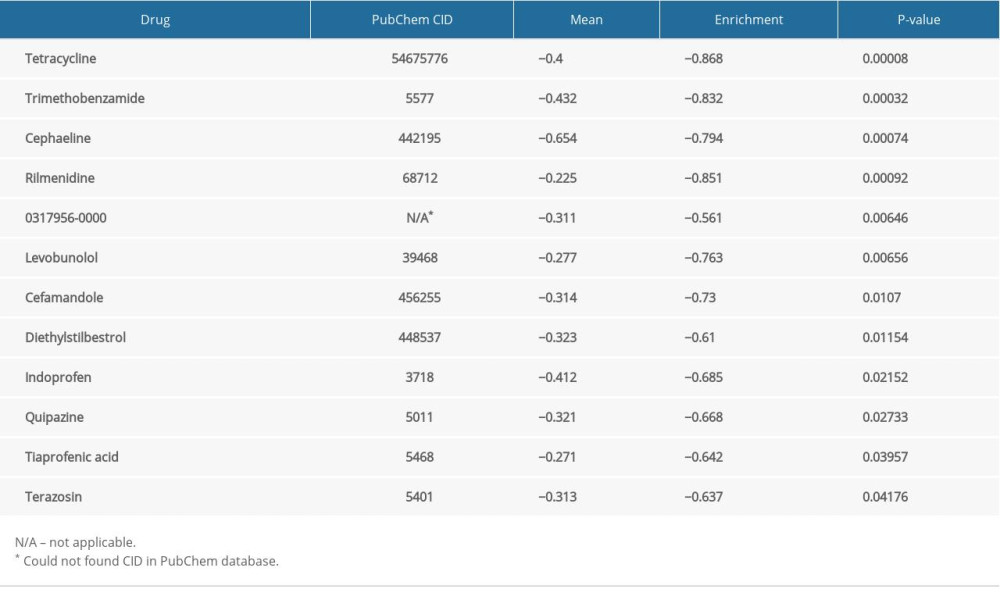 Table 5. Pharmacological targets and drug for ENO3.
Table 5. Pharmacological targets and drug for ENO3. Table 1. Demographic and clinical data for 438 colon cancer patients.
Table 1. Demographic and clinical data for 438 colon cancer patients. Table 2. Prognostic survival analysis of ENO family genes.
Table 2. Prognostic survival analysis of ENO family genes. Table 3. Grouping according to ENO genes.
Table 3. Grouping according to ENO genes. Table 4. Pharmacological targets and drug for ENO2.
Table 4. Pharmacological targets and drug for ENO2. Table 5. Pharmacological targets and drug for ENO3.
Table 5. Pharmacological targets and drug for ENO3. In Press
21 Mar 2024 : Meta-Analysis
Economic Evaluation of COVID-19 Screening Tests and Surveillance Strategies in Low-Income, Middle-Income, a...Med Sci Monit In Press; DOI: 10.12659/MSM.943863
10 Apr 2024 : Clinical Research
Predicting Acute Cardiovascular Complications in COVID-19: Insights from a Specialized Cardiac Referral Dep...Med Sci Monit In Press; DOI: 10.12659/MSM.942612
06 Mar 2024 : Clinical Research
Enhanced Surgical Outcomes of Popliteal Cyst Excision: A Retrospective Study Comparing Arthroscopic Debride...Med Sci Monit In Press; DOI: 10.12659/MSM.941102
06 Mar 2024 : Clinical Research
Prevalence and Variation of Medical Comorbidities in Oral Surgery Patients: A Retrospective Study at Jazan ...Med Sci Monit In Press; DOI: 10.12659/MSM.943884
Most Viewed Current Articles
17 Jan 2024 : Review article
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
14 Dec 2022 : Clinical Research
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387
01 Jan 2022 : Editorial
Editorial: Current Status of Oral Antiviral Drug Treatments for SARS-CoV-2 Infection in Non-Hospitalized Pa...DOI :10.12659/MSM.935952
Med Sci Monit 2022; 28:e935952








