21 November 2020: Database Analysis
Comprehensive Analysis of Alternative Splicing Signature in Gastric Cancer Prognosis Based on The Cancer Genome Atlas (TCGA) and SpliceSeq Databases
Xiaohu Cheng 1ACEG* , Xianghua Li 2ACE* , Yimei Gu 3AE , Lianbang Zhou 1AE , Jingjing Tang 1B , Xiang Dai 1B , Heng Jiang 1B , Yang Huang 1B , Yingfeng Zhang 1B , Tongtong Xu 1B , Zhining Liu 1AE* , Qihong Zhao 4AE*DOI: 10.12659/MSM.925772
Med Sci Monit 2020; 26:e925772
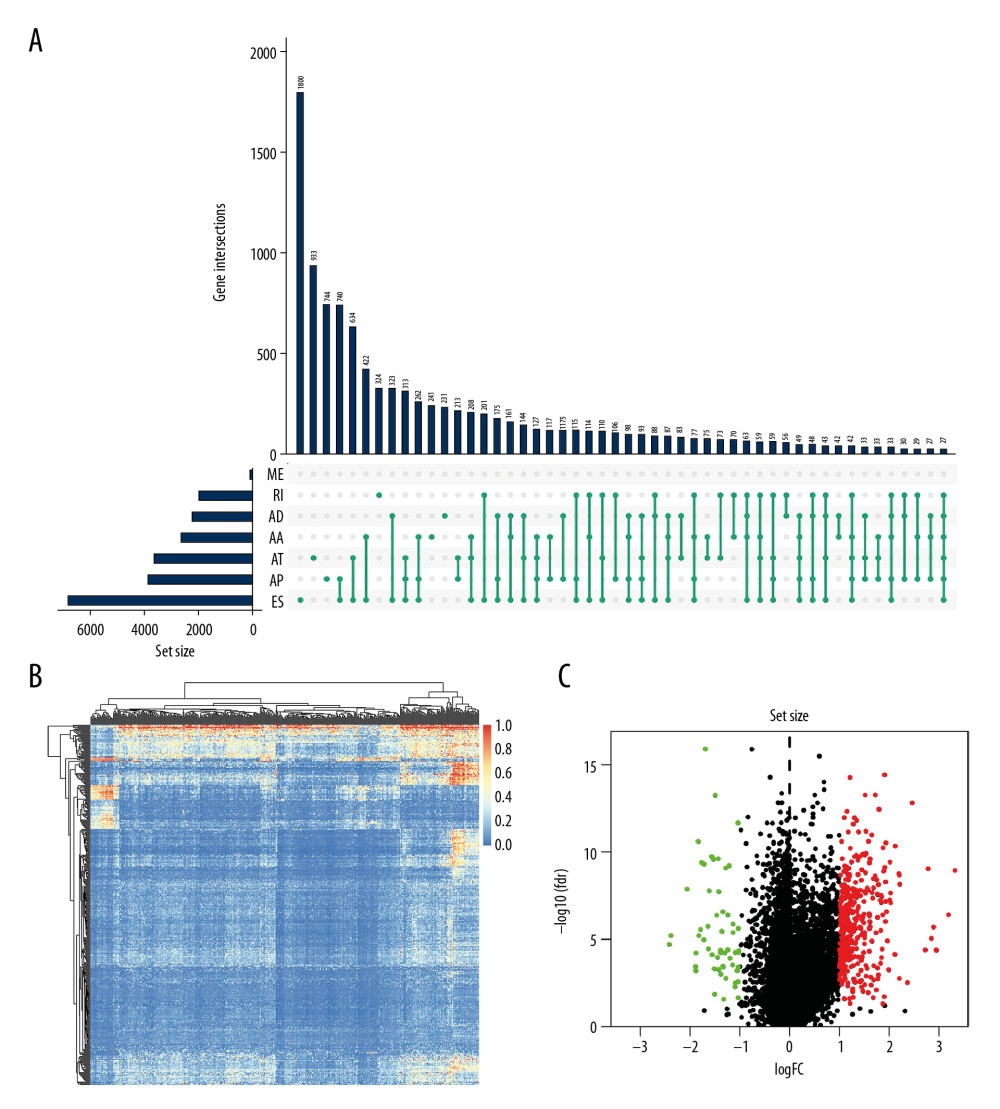
Figure 2 The landscape of aberrant alternative splicing events in GC cohort. (A) Upset plot of ASEs for the 7 different patterns, including AA as alternate acceptor site, AT as alternate terminator, ES as exon skip, AD as alternate donor site, AP as alternate promoter, ME as mutually exclusive exons, and RI as retained intron in the GC. (B) The expression heatmap of differentially expressed alternative splicing events (DEAS). (C) Volcano plots of the distribution of DEAS in the GC dataset. Red dots represent upregulated alternative splicing events whereas green dots represent downregulated ones.


